Our first international workshop with from ltr; Anders Hobæk (NIVA), Cessa Rauch & Jon Kongsrud (UMB), Tone Falkenhaug (project leader, IMR), Alexandra Savchenko & Rony Huys (NHM), photo by Alexandra Savchenko
During the last week of September, HYPCOP organized its last and crucial workshop for finishing the project. We invited international collaborators Prof. Dr. Rony Huys and Dr. Alexandra Savchenko from the Natural History Museum in London. Prof. Dr. Huys is a well-known copepod taxonomist and crustacean researcher and published a multitude of species descriptions and books including key identification guides. We were very happy to hear he had time to come and travel to Bergen, paying us a visit while also helping us with species identifications of the many, many copepods we had collected during the two years of our project.
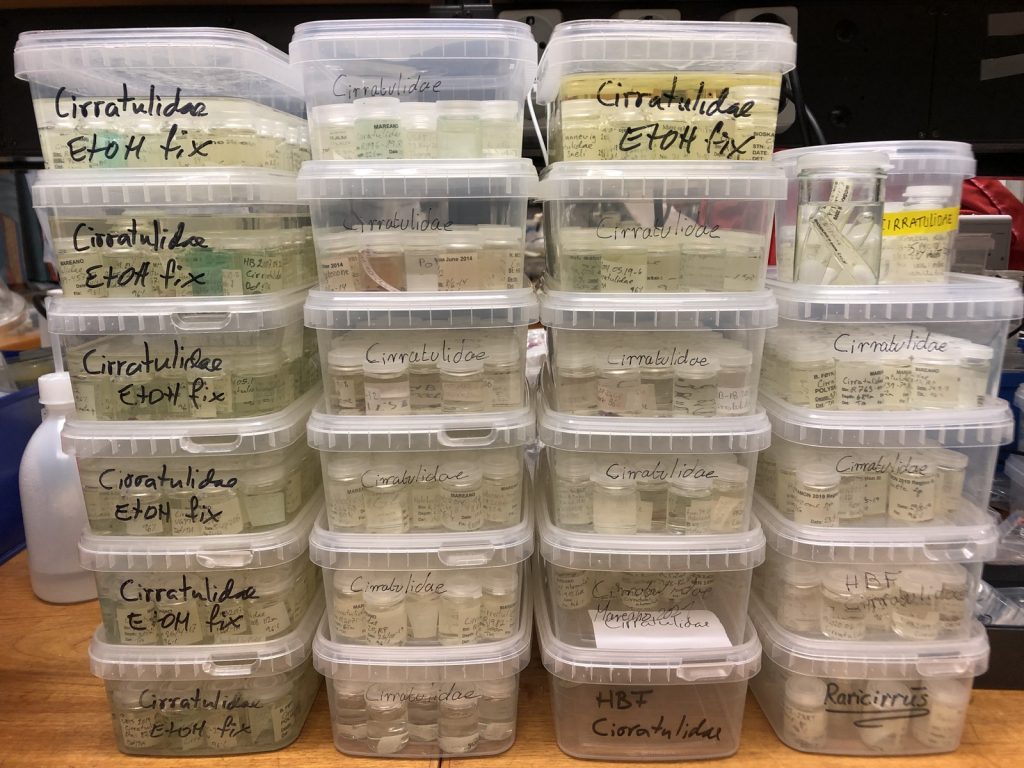
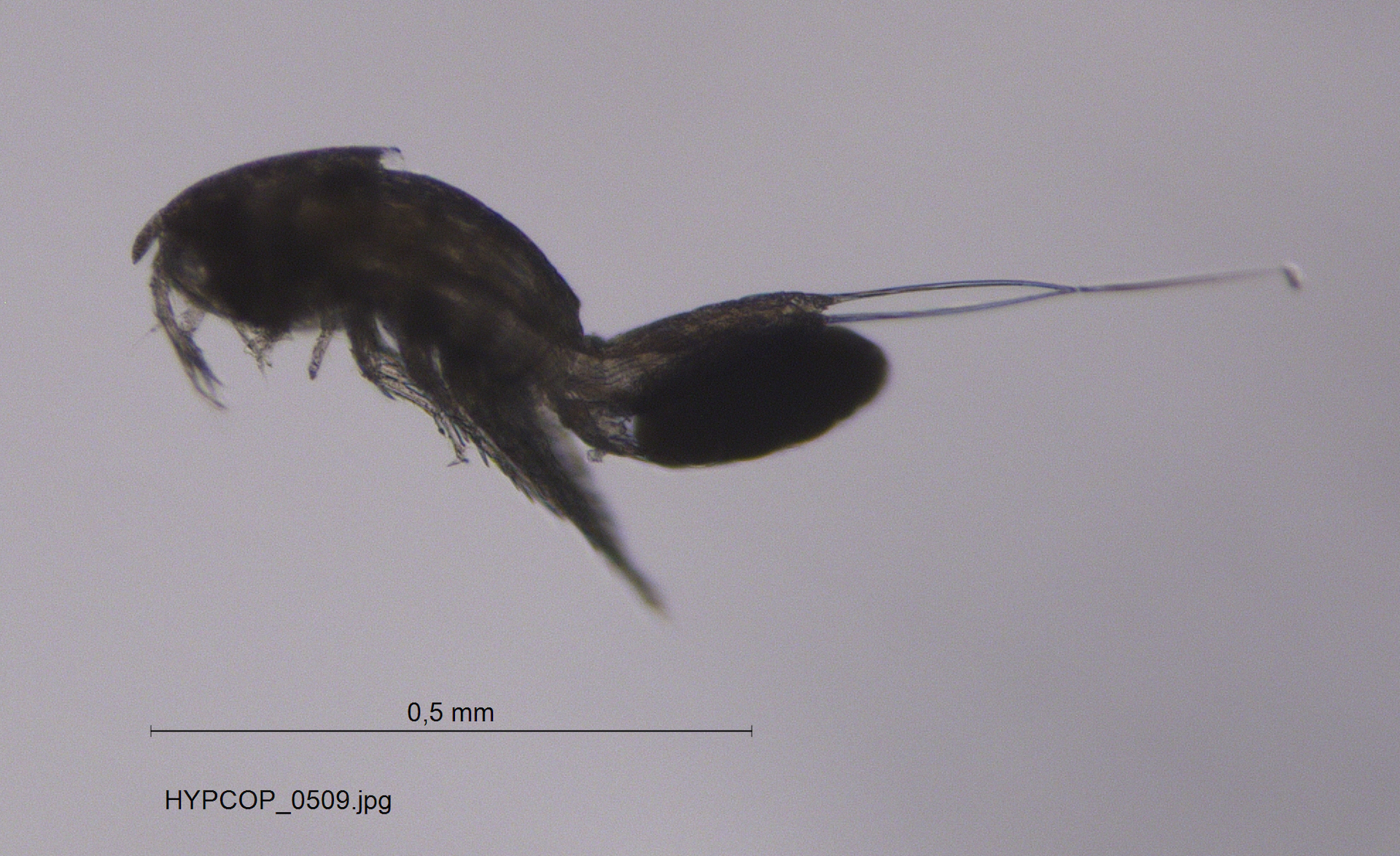
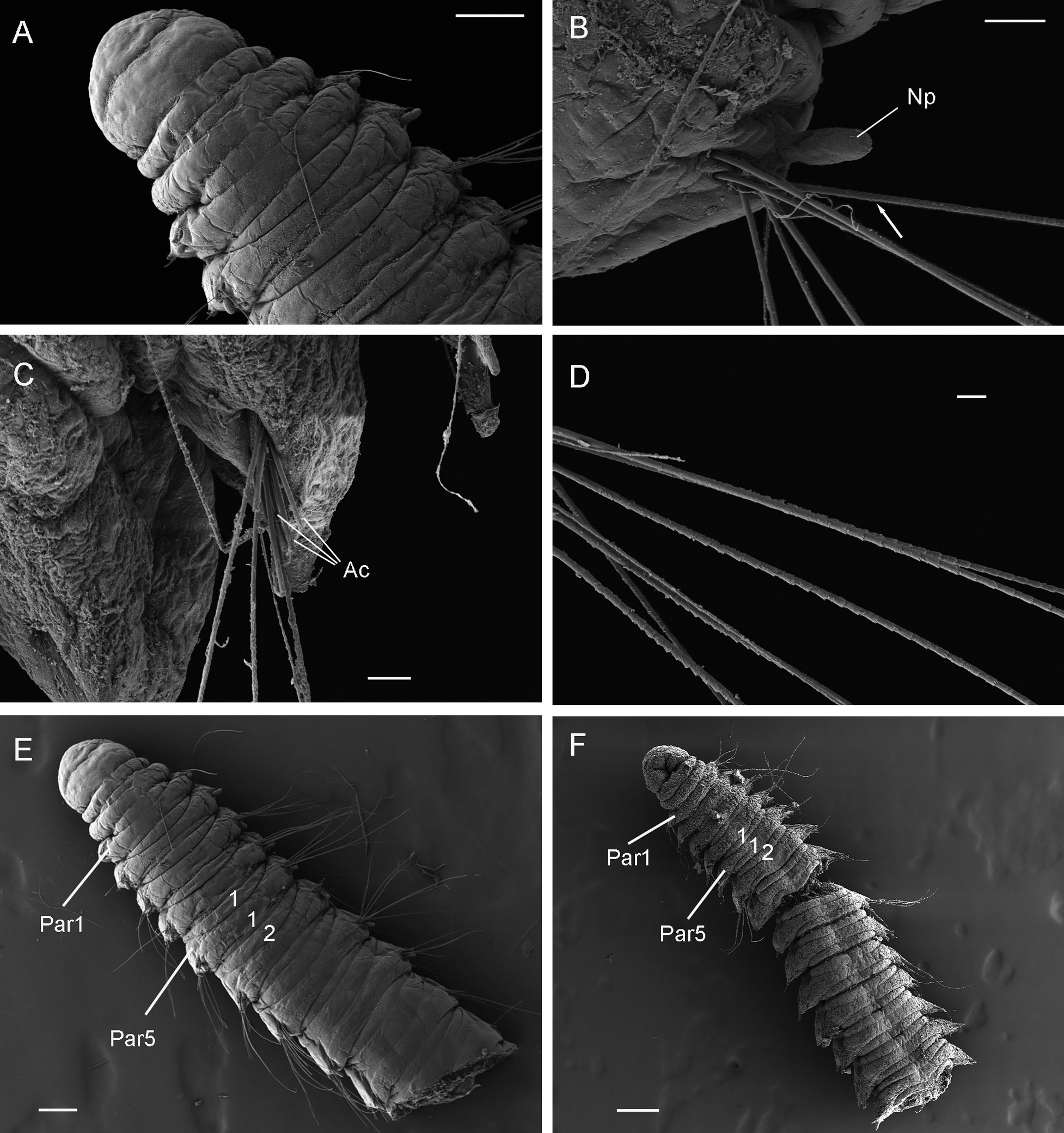
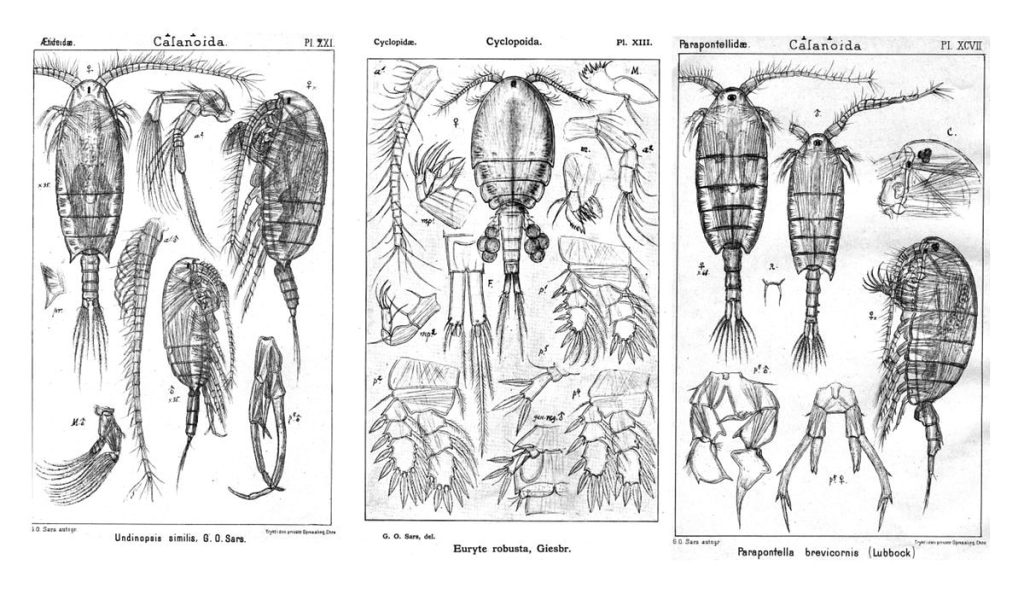
During the two years of the HYPCOP project we collected around 600 specimens from different localities all over Norway, including shallow coastal waters and the deeper parts of the mid-Atlantic Ridge (Loki’s Castle field of active hydrothermal vents). From all those specimens we extracted DNA from the soft tissue of the animal. Therefore, keeping the hard exoskeletons, for morphological identification downstream. This is the most time consuming and challenging part. The species can sometimes only be identified based on minuscule differences in the appearance of its legs. Besides, one needs good taxonomic competence to assign these differences to the thousands of marine benthic copepods species. And this is where the HYPCOP team needed help.
HYPCOP started in May 2020, when a lot of countries, including Norway, were in a lockdown and international travel was difficult or even impossible. Therefore, it was problematic for HYPCOP to invite international researchers for most of the time. Thus, we focused mostly on extracting DNA from our collected specimens and building up a barcode library. But what was missing was the nomenclature of the bulk of the specimens. When finally, our first international researchers could come and have a look at our specimens, it turned out to be an enormous task. With the help of Prof. Dr. Huys and Dr. Savchenko we managed now to have almost 300 assigned names to our DNA library of 500 specimens. Quite a few of those are new species and even new genera.

Kickoff of the workshop, which would take place at Marine Biological Station Espegrend for the duration of a week, photo by Alexandra Savchenko
Rony and Alexandra arrived Sunday evening in Bergen together with project leader Tone Falkenhaug and project technician Cessa. We were stationed at the Espegrend marine biological station in Bergen for the entirety of the week. It was for Tone and Cessa the first time they would finally meet Rony and Alexandra in person, after many months of digital communication. It was a nice relaxing first evening. The next day Anders Hobæk from NIVA and Jon Kongsrud from the UiB joined and we started off the week with a presentation overview of the project.
The overview informed everyone about the program of the week and the state of the art of the project. With the DNA barcode library, we managed to construct a COI phylogenetic tree. Some of the larger clades were already identified down to species level, but many more species names were missing from the smaller clades. It was up to us that week together with Rony and Alexandra to identify these last cases.

Alexandra onboard research vessel Emiliana, photo by Tone Falkenhaug
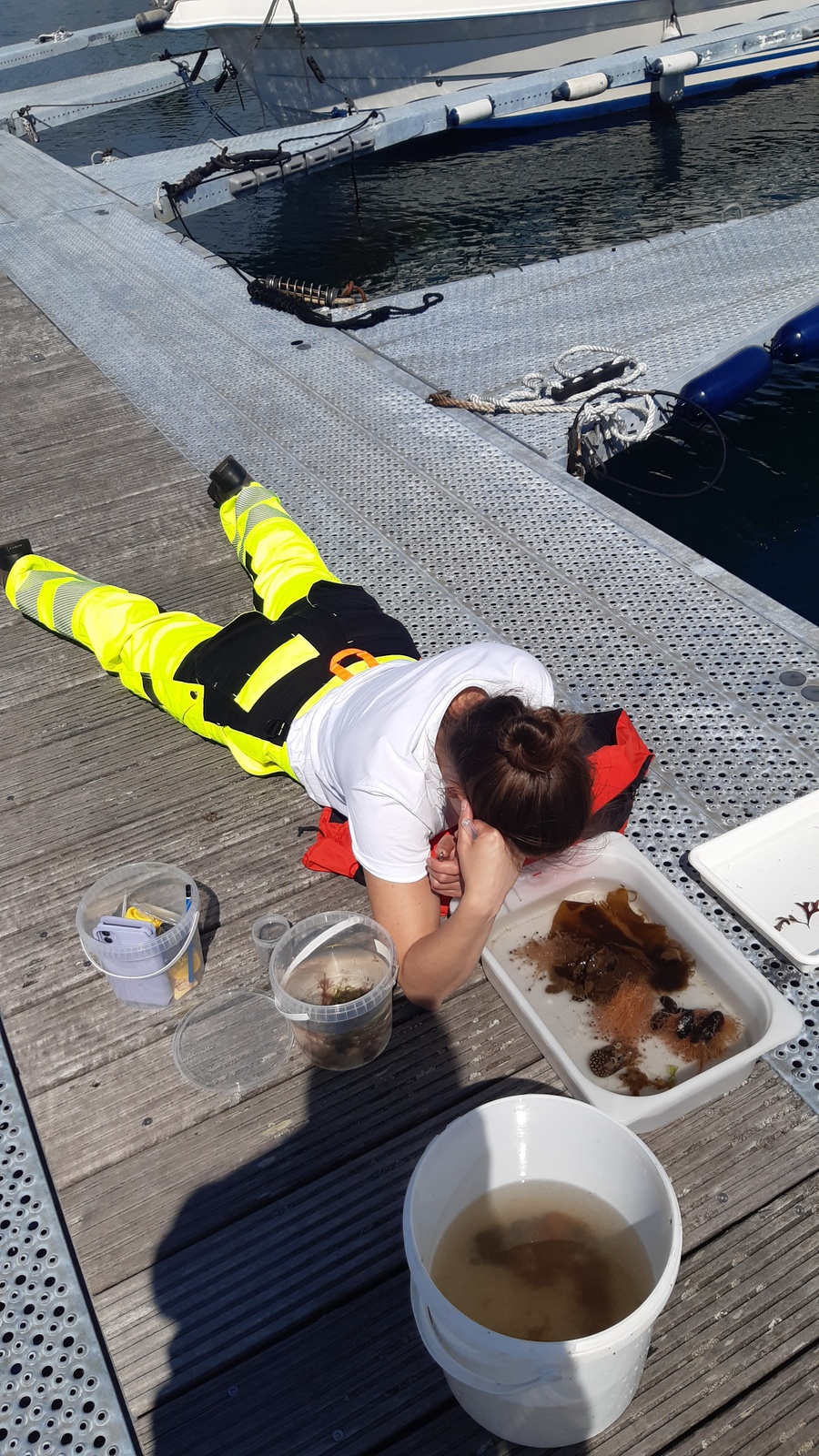
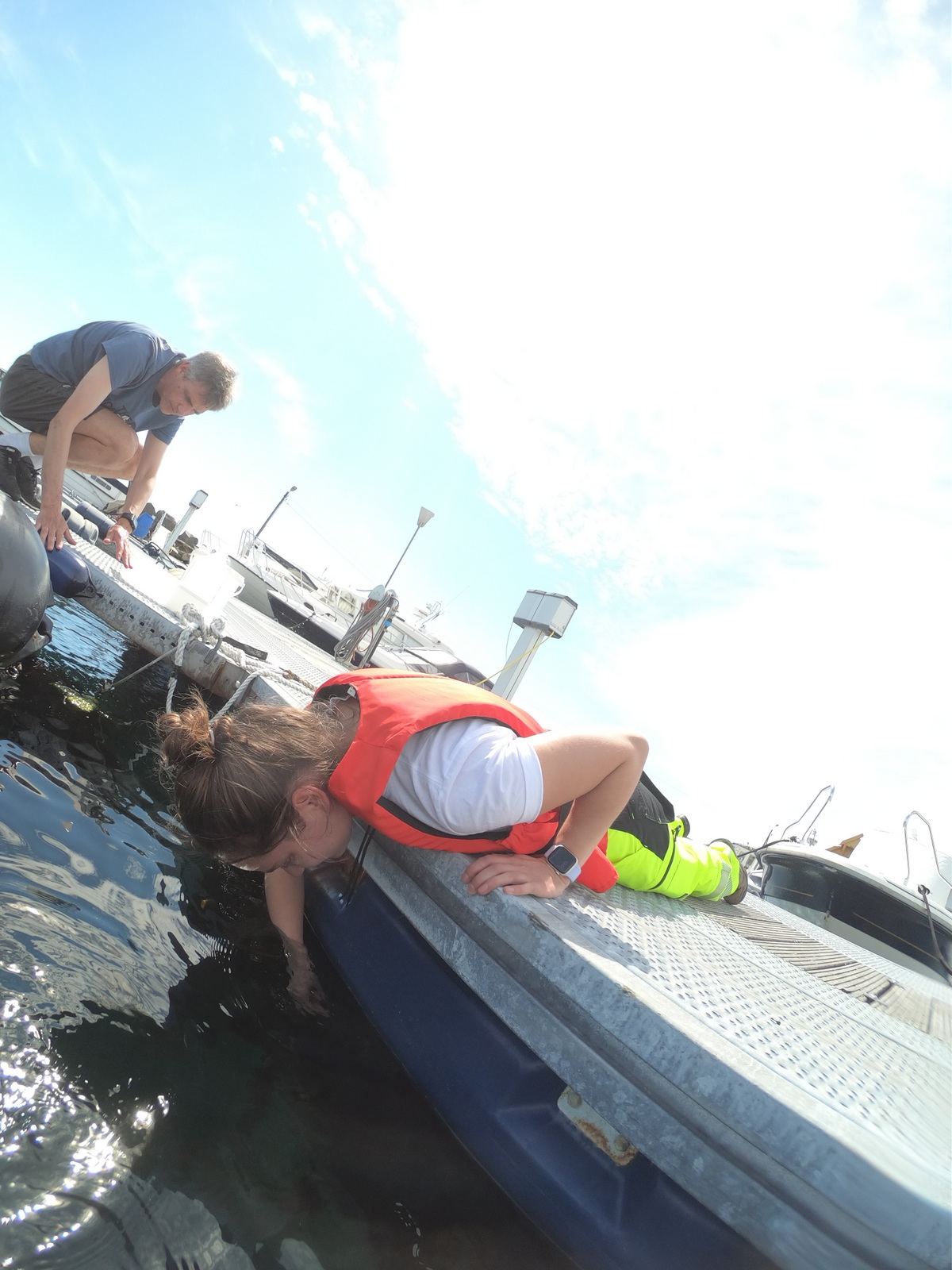
We also had one day of fieldwork planned, to have us work also with some fresh material. This we did with help of research vessel Emiliana and the Beyer’s sled. Both stationed at Espegrend Marine Biological station. We tried to pick out a nice and dry day for going out with the boat and that happened to be in the mid of the week. We went a little bit outside of the Biological Station, with a depth of around 90 – 120m. The Beyer’s sled is an epibenthic sampler, it is called a sled for its form. We got many fresh samples, but due the net being a little large in its mesh size, we did not get as many small species as we liked.
-

-
Anders helping with the Beyer’s sled on board of r/v Emiliana, photo by Cessa Rauch
-

-
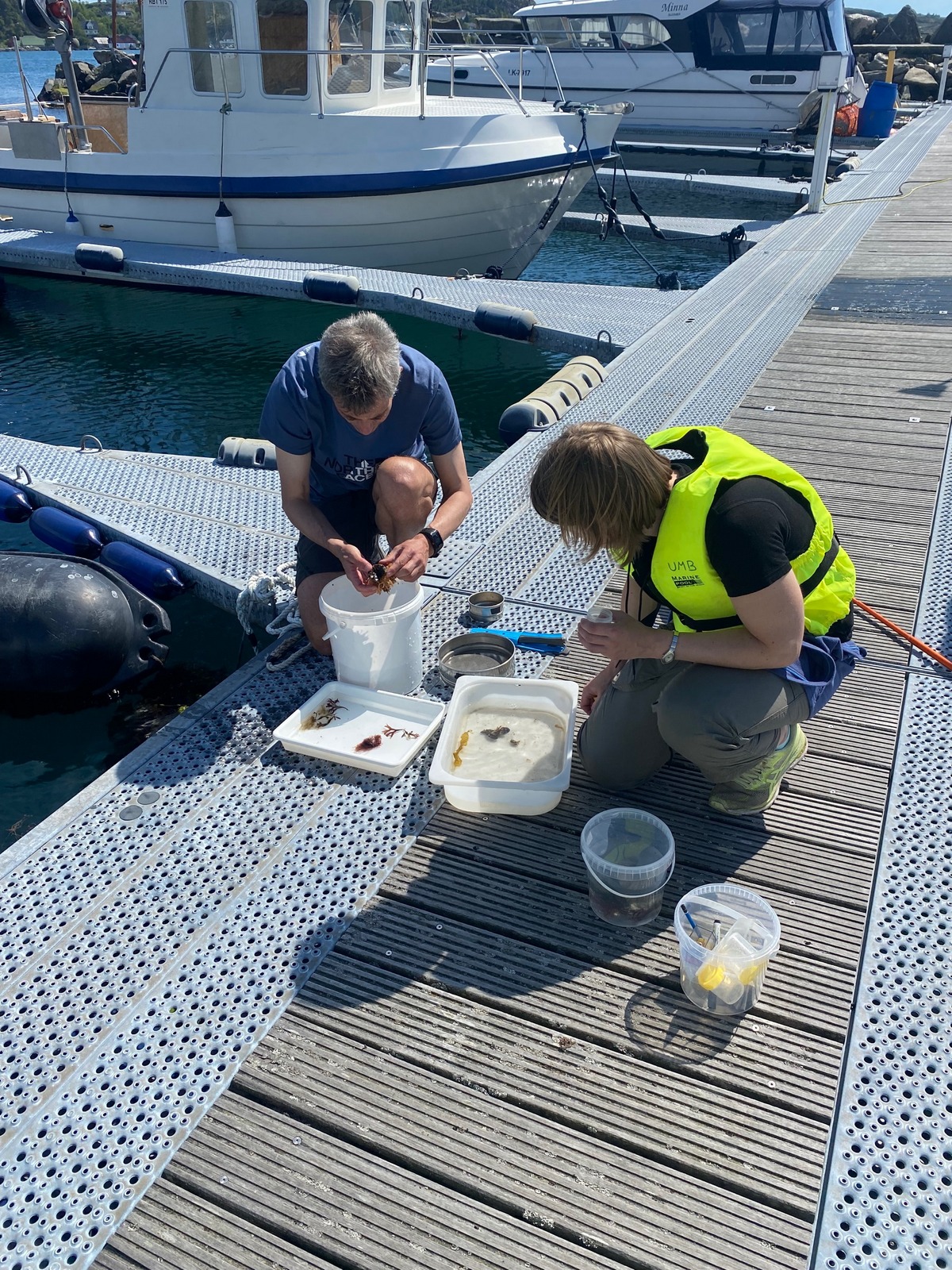
Alexandra and Rony sieving the catch of the Beyer’s sled earlier that day, photo by Cessa Rauch
-

-
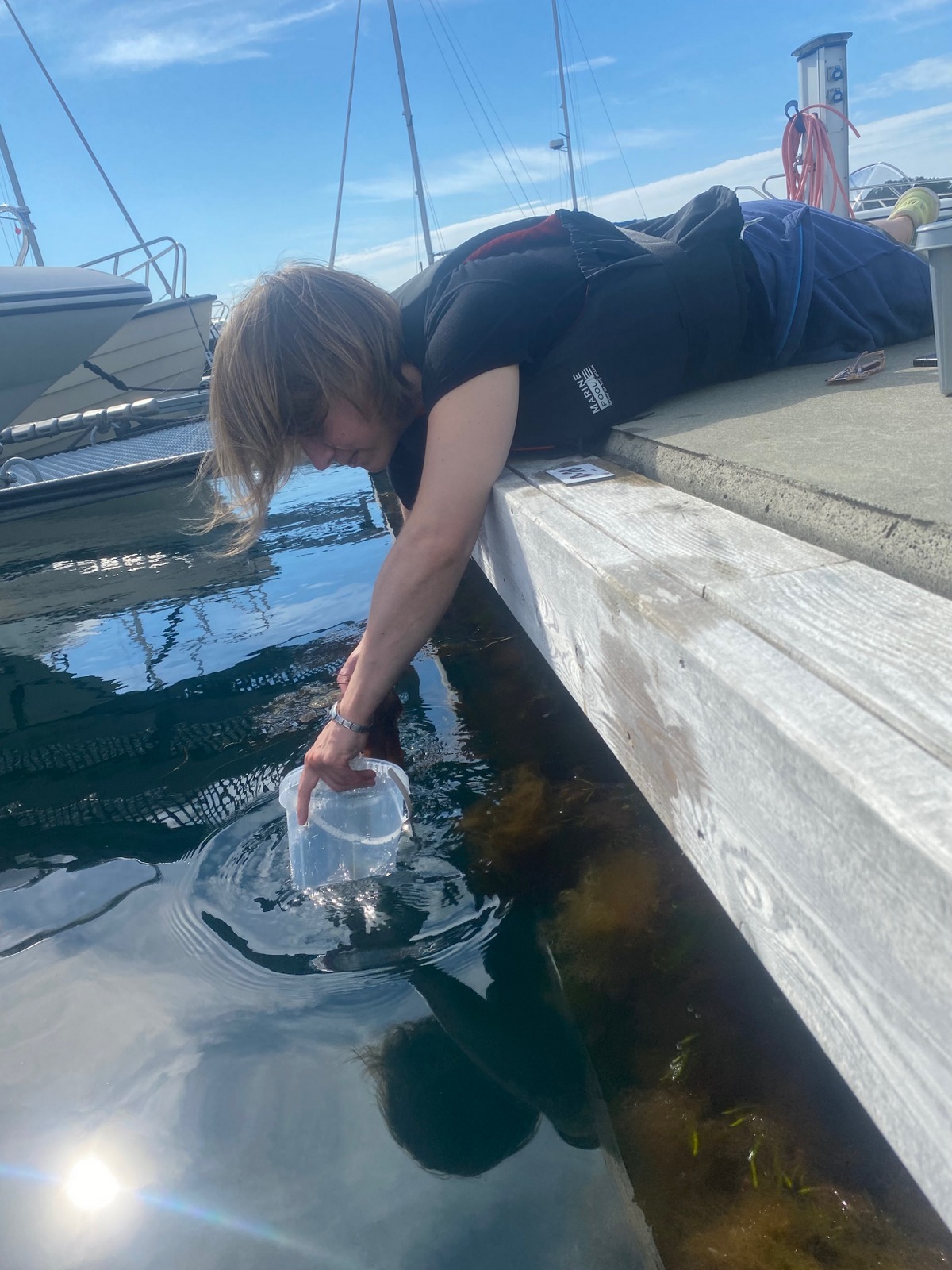
Example of a DIY light trap; a bottle with a funnel as opening and a small LED light on the bottom, photo by Cessa Rauch
Therefore, we also tried another sampling method with help of Anders; he had brought with him a light trap. Light traps are very easy to DIY with a bottle and inverted bottle opening, like a funnel, and a small led light on the bottom. You install the trap in the water overnight; the little led light attracts a lot of small hyperbenthic and planktonic (and some bigger) species.
-

-
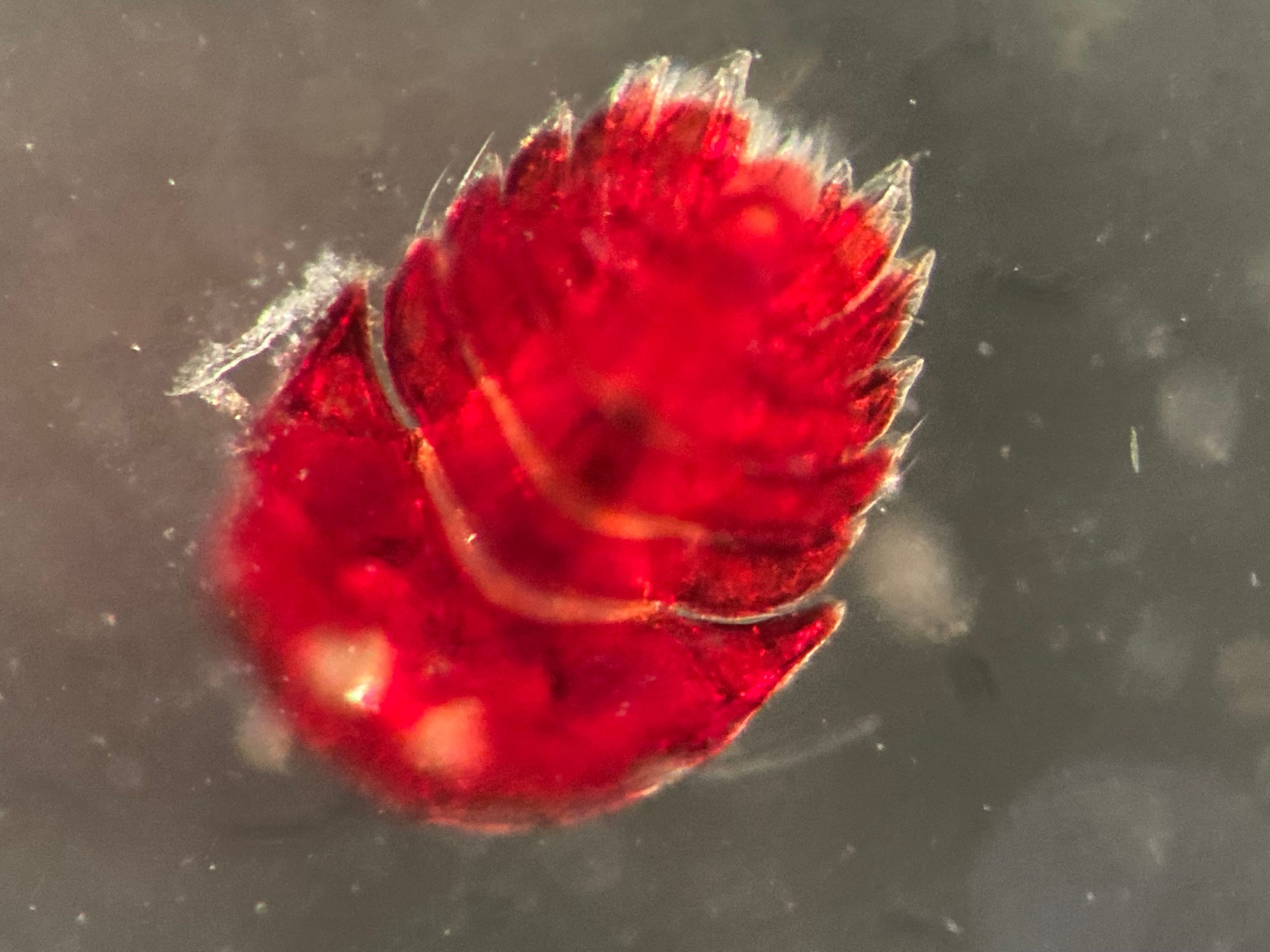
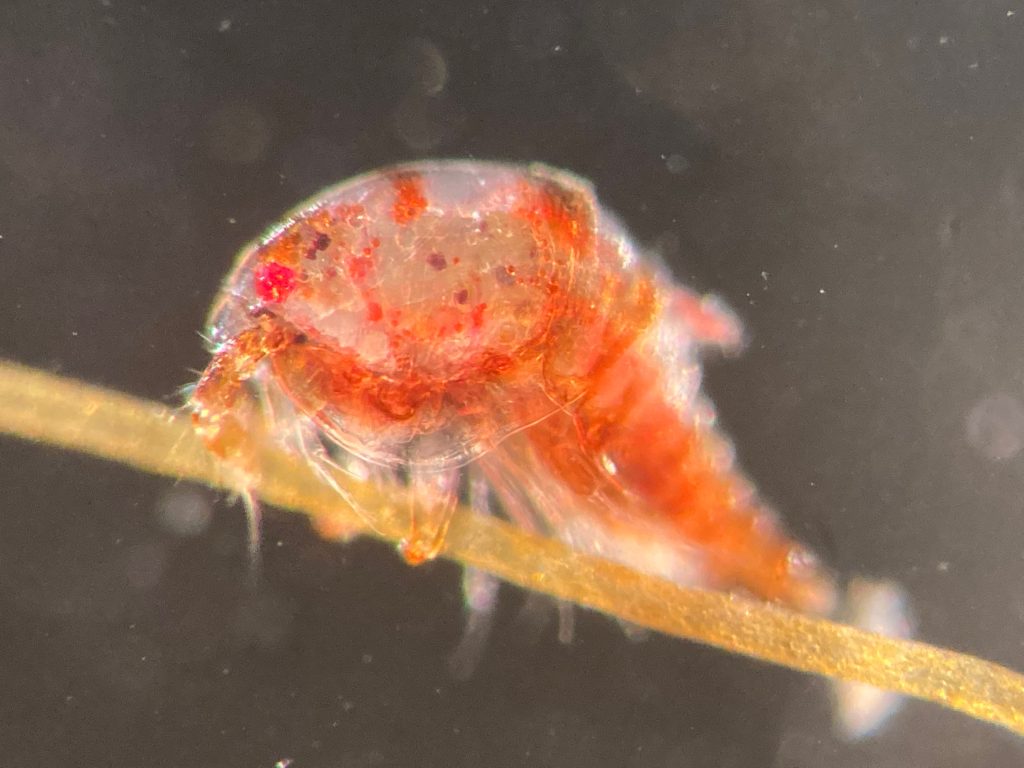
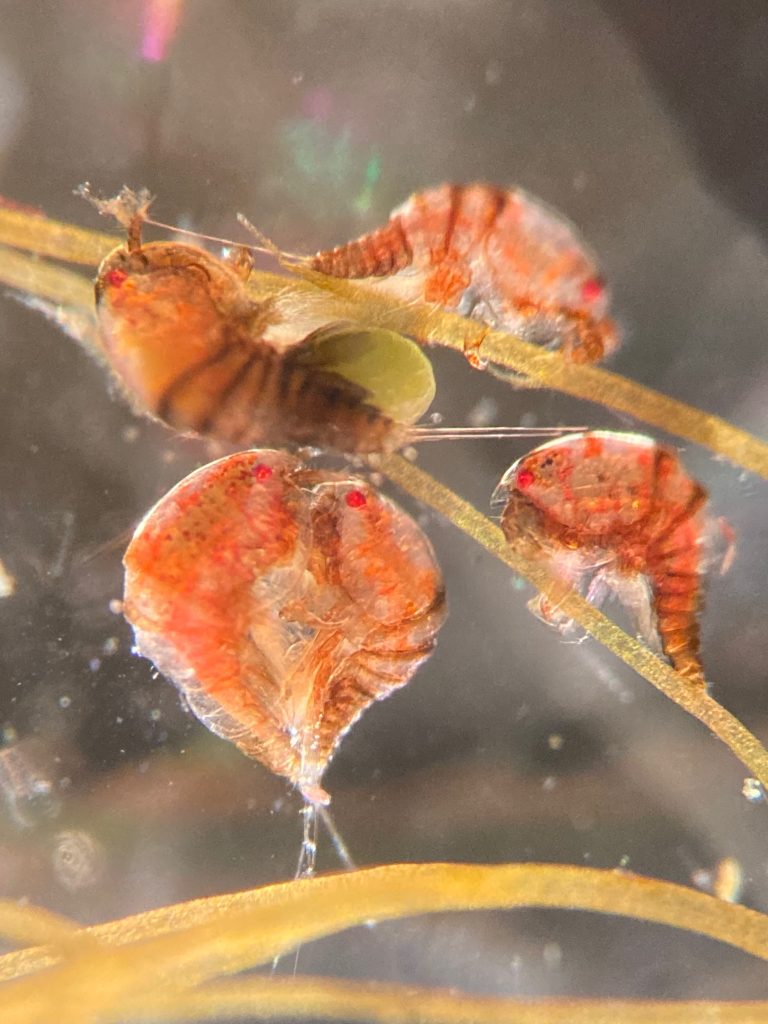
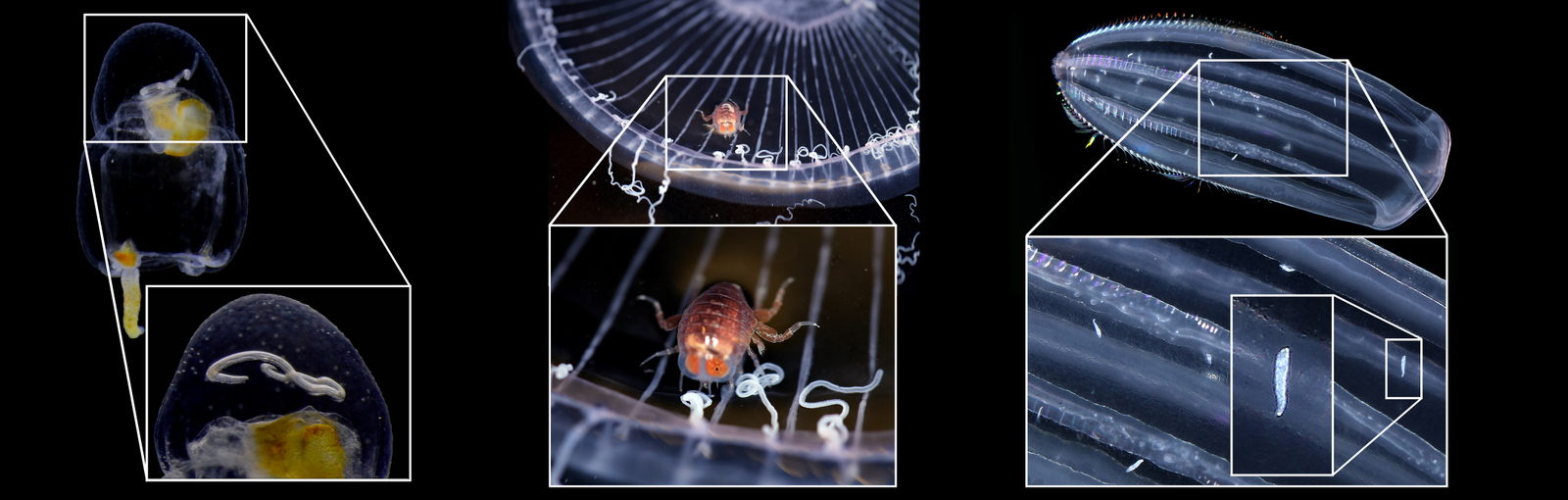
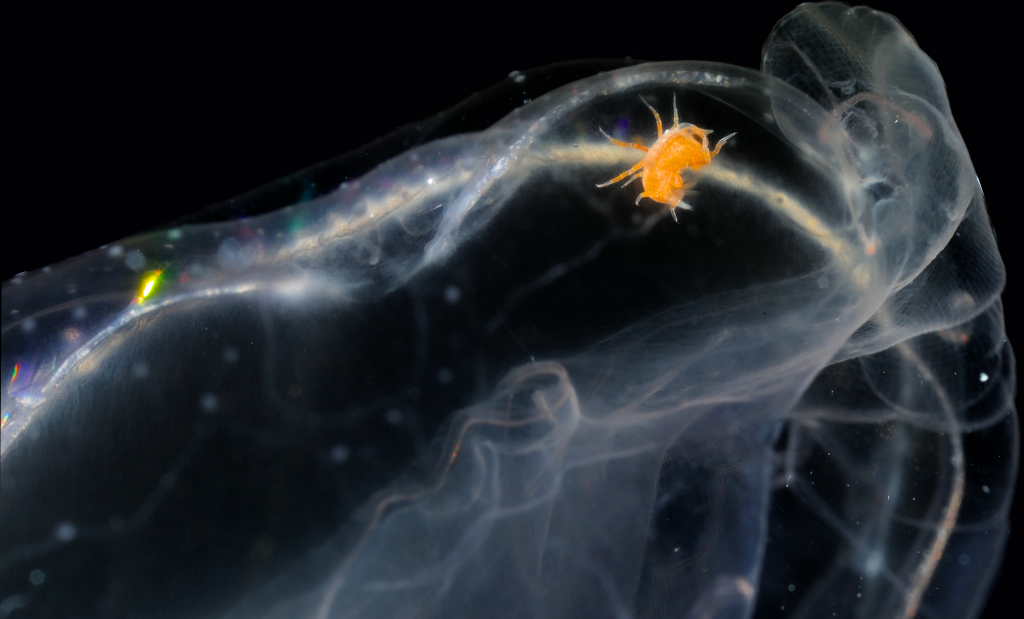
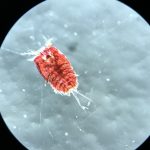
The focus of the week was working on fixed material, but it is much more fun to work with freshly caught specimens, that still have their beautiful colors, photos by Alexandra Savchenko
-

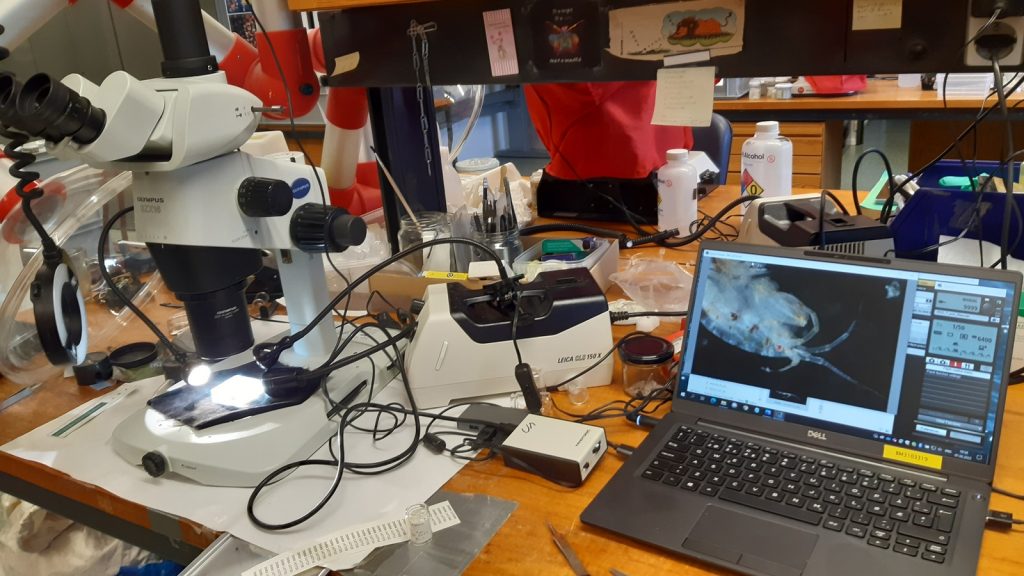
Everyone working hard at the Marine Biological Station Espegrend, assigning species names to specimens, photo by Cessa Rauch
The entirety of the week consisted of many hours working at the microscope, going through literature, dissecting specimens, and assigning species names to the specimens. Eventually with help of Rony and Alexandra, we managed to assign 298 scientific names to 702 specimens in our collection. From those specimens, we extracted DNA from 593 specimens and produced a DNA library, which we uploaded to the BOLDSYSTEMS (Barcode of Life Data System). This library also has all the metadata of our specimens, such as location, depth, size, and pictures of the specimens (either life, fixed and in some cases parts). And it will be publicly available at the end of the HYPCOP project.
The week was demanding but very rewarding and we got many specimens identified, with even a few new species and genera to Norway and possibly new to science; all thanks to the hard work and help of Rony and Alexandra. We therefore also would like to take this opportunity to thank them again for their time and efforts in helping the HYPCOP project move forward! Until next time.

Rony Huys and Alexandra Savchenko helping the HYPCOP project move forward, photo by Tone Falkenhaug
– Cessa