The project of studying hyperbenthic copepods (HYPCOP) is unique in multiple ways; we study a very unknown group of marine copepod species with very little taxonomic knowledge available here, in Norway. It is challenging as there are more than 700 species described, and possibly more. With pandemic lockdowns, it was hard to have international specialists come and help us, so we had to rely on resources available locally. With so many institutes involved from different corners of Norway, it was not always easy to meet up physically to work on our collection. Hence, when it happens, it is a memorable event, and valuable progress for the project is made.
We have Tone Falkenhaug as the project leader, situated at the Institute of Marine Research in Flødevigen (IMR), than we have our collaborator from Norwegian Institute for Water Research (NIVA), Anders Hobæk and the three technicians at the department of natural history from the University of Bergen. The year before we all got together in Flødevigen, so for 2021 we decided that it would be Bergen to have another workshop.
A year into our project we managed to build up a substantial collection of benthic copepods; currently we have around 460 registered specimens, and 195 off those are barcoded with two different DNA markers, mitochondrial (COI) and ribosomal (16S). What keeps ahead of us is the monster task of working through our specimens to label the DNA barcodes with morphological identifications. It means many hours of very precise work with the finest needles, while sitting at the microscope.
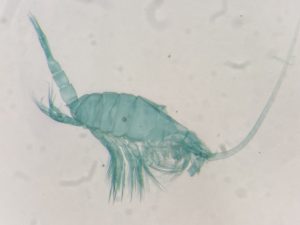
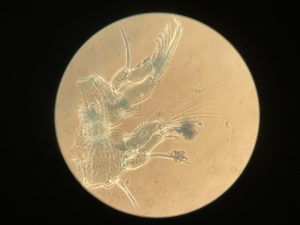
- Copepods being stained and dissected for morphological identification
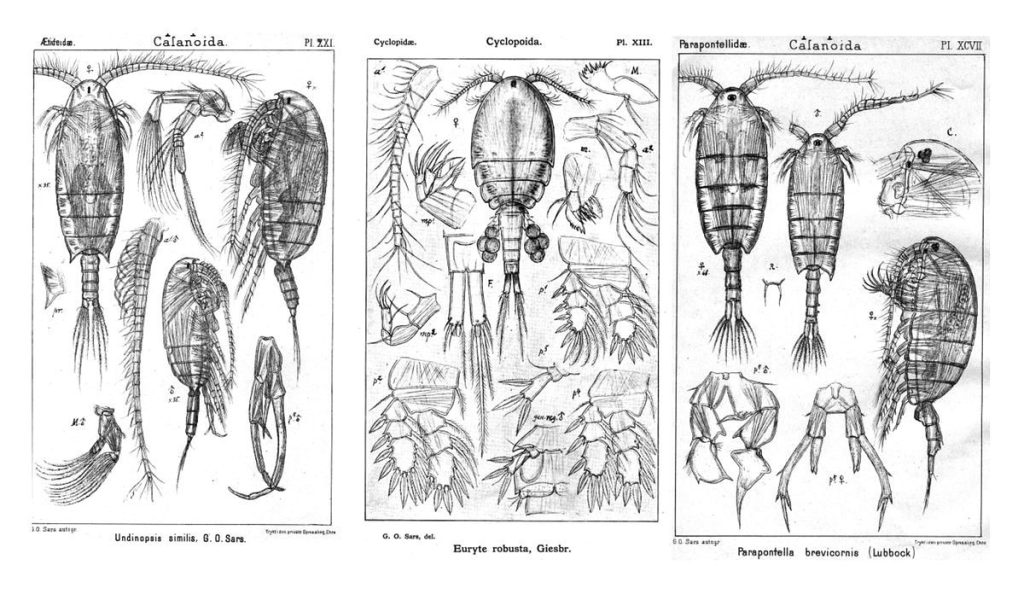
During our workshop in Bergen we got together to work through one of the copepod family trees we generated from their DNA:
Anders Hobæk is a taxonomist with many years of experience dissecting copepods, and together we went through the samples one by one. It is very satisfying to be able to identify a specimen and get the to the same species level as the DNA barcode. There are multiple reasons as why we choose to identify species based on morphology.
Not all species are easy to barcode, as copepods, especially the benthic ones, are often so extremely tiny; it is difficult to get good quality DNA extracted from them.
The small quantities of copepod DNA goes hand in hand with greater risk of contamination of other surrounding DNA, especially if you work with more general markers. Besides, even if we have the DNA barcode, not all copepod DNA is identified as such, which means that even with the right DNA, when running it through the database, it tells us that we have fly DNA, to give an example. Last but not least, in a lot of cases, we were not able to get good DNA sequences from the copepod extracts, so the only option is identifying them morphologically, by dissecting the animals and with help of literature identify the right genus, or even better, the species.
Our next workshop shall take place again in Flødevigen, in the meantime we keep you updated about our planet of the copepods.
Follow us for more copepod content @planetcopepod, see you there!
-Cessa