Monthly Archives: December 2019
Guest researcher: Eric
Eric, from the Federal University of ABC, visited the University Museum in November. We asked him about his time in Bergen examining some of the least common species of siphonophores in the collections and this is what he told us:
My name is Eric Nishiyama, and I am a PhD student from Brazil. The main focus of my research is the taxonomy and systematics of siphonophores, a peculiar group of hydrozoans (Cnidaria, Medusozoa) notorious for their colonial organization, being composed of several units called zooids. Each zooid has a specific function within the colony (such as locomotion, defense or reproduction) and cannot survive on its own.

Fig_1. I had the opportunity to examine both ethanol- and formalin-fixed material from the museum. For morphological analyses, specimens preserved in formalin are preferable because ethanol-fixed individuals are usually severely deformed due to shrinkage.
Understanding how zooids evolved could provide major insights on the evolution of coloniality, which is why I am looking at the morphology of the different types of zooids. In this sense, siphonophore specimens available at museum collections provide valuable information for visiting researchers such as myself.
During my short stay at the University Museum of Bergen in November, I was able to examine a few siphonophore samples deposited at the museum’s collections. By examining the specimens under a stereomicroscope, and using photography and image processing tools, I was able to gather a lot of information on the morphology of several species.
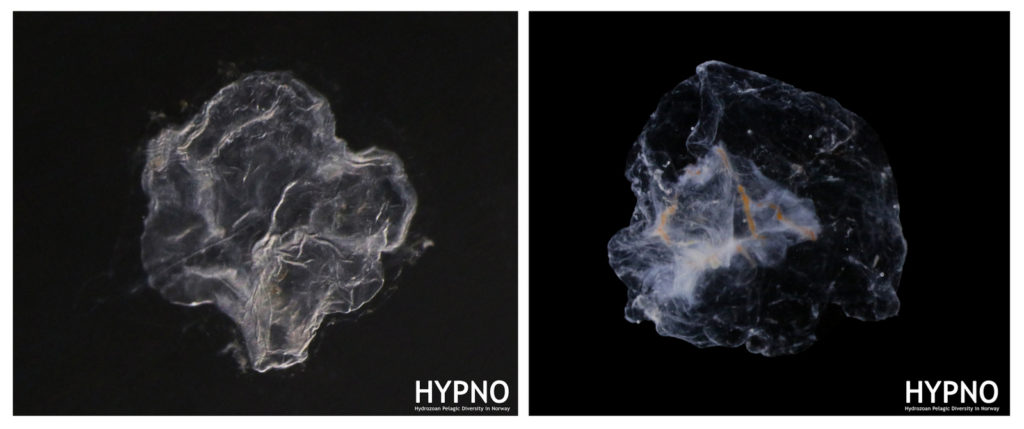
Fig_2. Documenting the morphology of the nectophores of Rudjakovia plicata (left) and Marrus or-thocanna (right) was particularly interesting because these species are not commonly found in museum collections.

Fig_3. Other ‘unusual’ siphonophores that I was able to examine were Crystallophyes amygdalina (left) and Heteropyramis maculata (right).
The data obtained will allow me to score morphological characters for a phylogenetic analysis of the whole group, and hopefully will help me revise the group’s taxonomy.
– Eric
CT scan all the things!
Oslo Natural History Museum 02.12.2019 – 06.12.2019
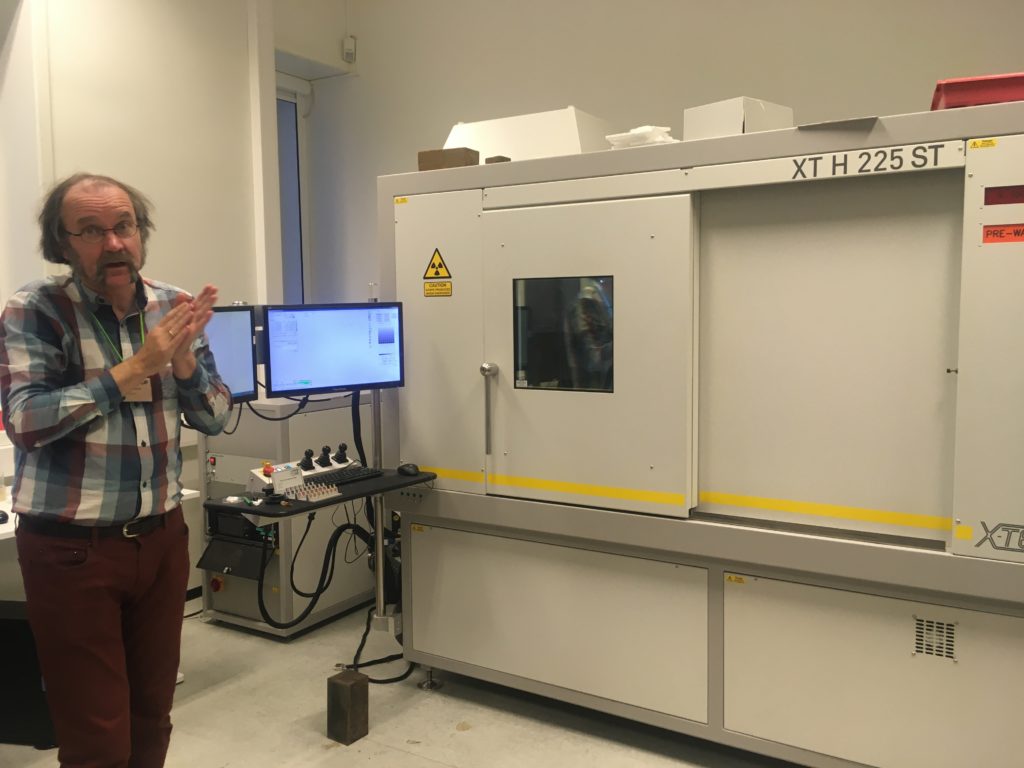
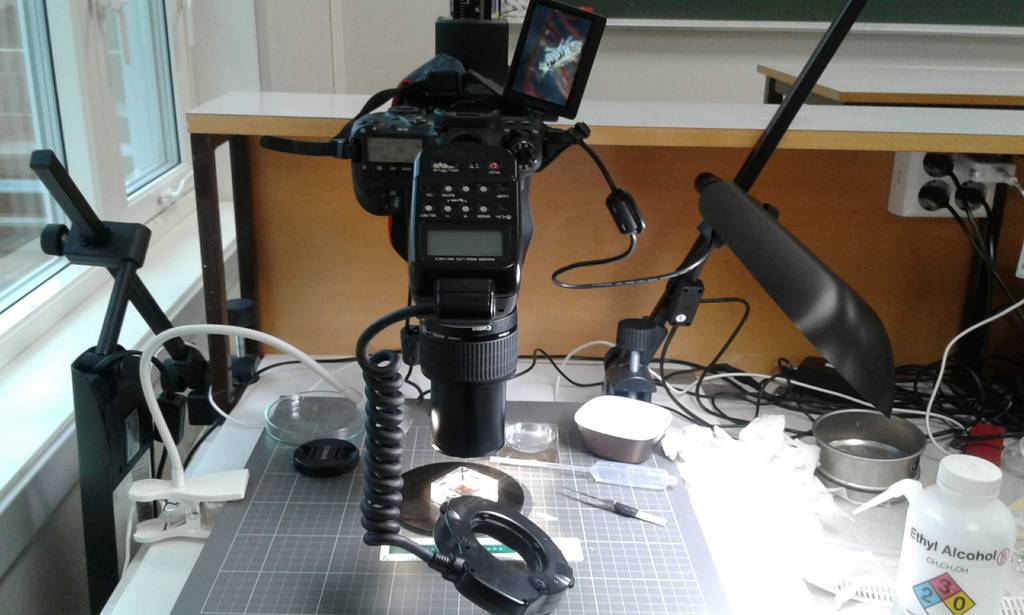
From December 2 till December 6 Justine Siegwald, Manuel Malaquias and I (Cessa) attended a CT scanning course in Oslo at the Natural History Museum.
The course was organized by the Research School in Biosystematics (ForBio) and Transmitting Science.
The three of us all work on mollusks and one of the main reasons to deepen ourselves in CT scanning techniques is because of studying the internal anatomy of these often small and delicate specimens.
Species descriptions are not only based on external morphology and/or DNA barcoding, a lot of the species differences are small details within the internal structures. Like the reproductive organs and digestive system (e.g. radular teeth) of the animals. This is very laborious work, and can be challenging especially with the smaller specimens of only a few millimeters long. Besides some species are difficult to collect and can be a rare collection item of the museum. Once you cut these animals to study them, there is no turning back and the specimen is basically destroyed. Being able to see the insides of the animal without touching it would therefore be ideal. CT scanning comes very close to that and with powerful X-ray we can almost see every detail of the insides of the animals without the need to cut them open
However, how easy as this sounds, CT scanning soft tissue comes with some challenges. Soft tissue means that the x-ray contrast is often very low. Even with modern good x-ray detectors it is still difficult to detect the different internal structures. Therefore, during the course, we were taught how to artificially increase the contrast of the tissue by staining the specimens before mounting them in the CT scan machine.
Coating a specimen with a metal (e.g. gold or platinum) is only useful if you want to see external details of the study object, coating will not help revealing internal structures. For radio contrasting the internal anatomy, you can stain specimens with high density liquids like iodine to enhance the x-ray contrast. So, we went for this option and left our specimens in iodine ethanol solution overnight
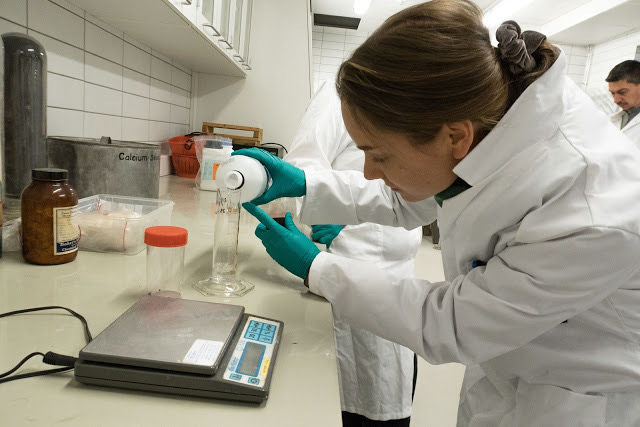
- Preparing the iodine solution
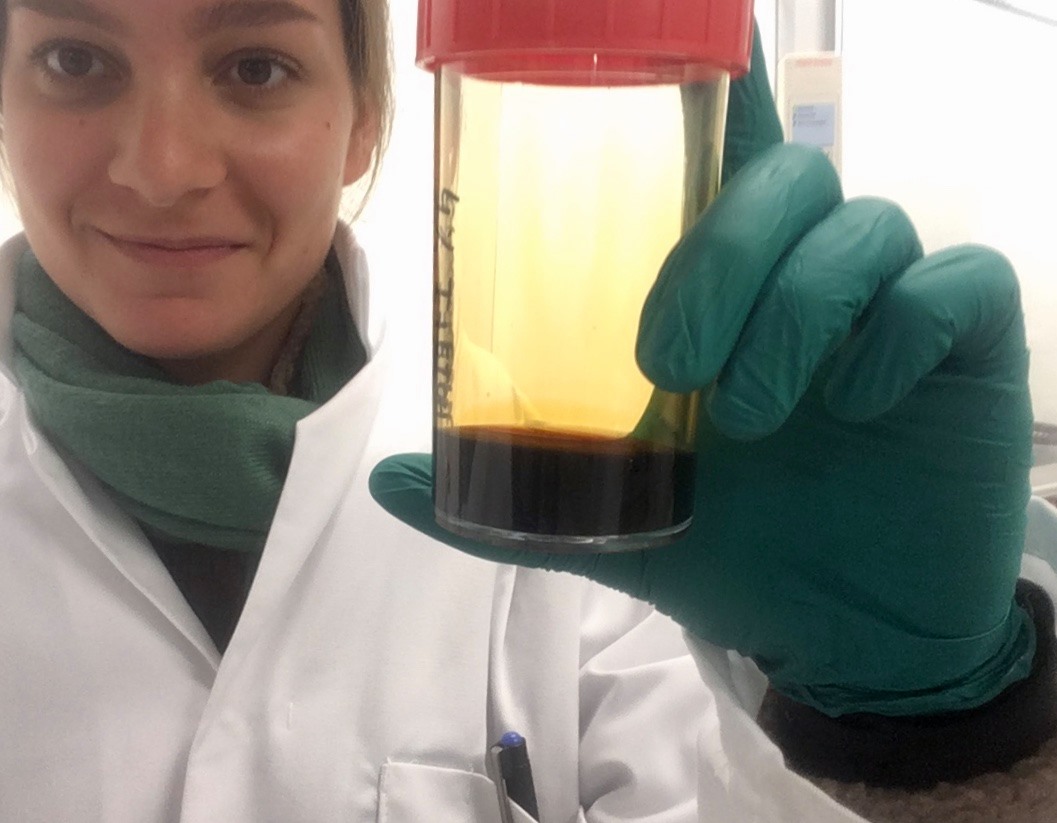
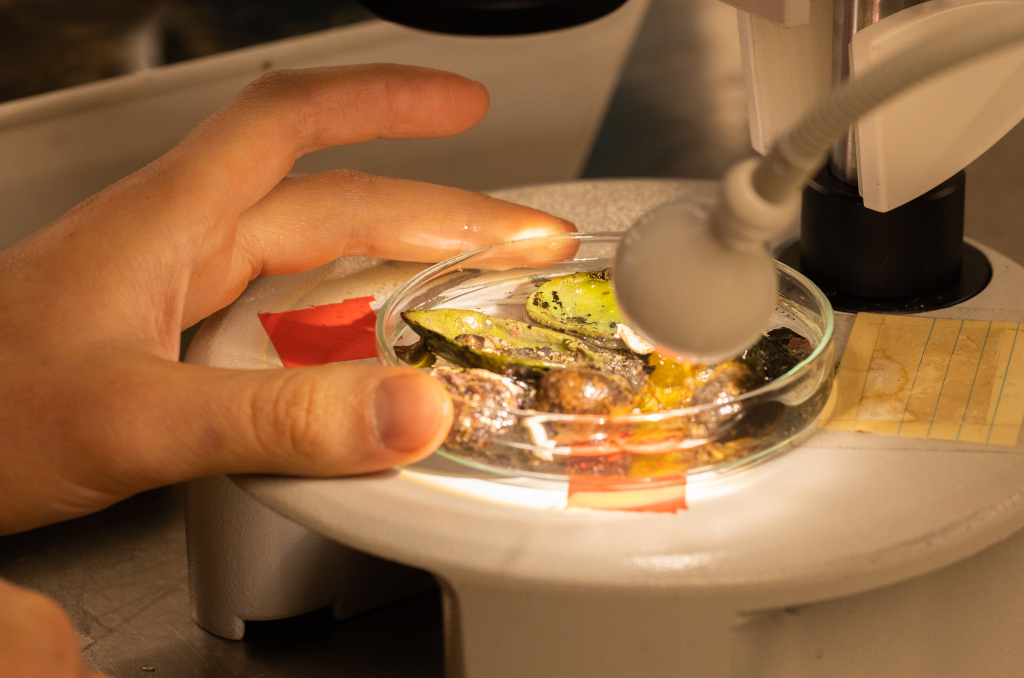
- Sea slug in iodine solution
After the staining process, we needed the wash out any extra iodine before mounting the specimen into the CT scan. A good scan is not simply pressing the button of the machine; there are a bunch of settings that can be adjusted accordingly (e.g. tube voltage, tube current, filament current, spot size, exposure time, magnification, shading correction, etc.). The scan itself can take hours and file sizes of 24GB for a single object are not uncommon, which means you need a powerful computer with decent software to process this information. Part of the course was also about how to visualize and analyze the files of the CT scan with software like Avizo
Video 1. Software Avizo has a user-friendly interface for analyzing all sorts of CT scanning files.
The week gave us some very promising first results (se video below), and also new insights about how to increase the X-ray contrast of our sea slug samples (lots of hours of staining and very short washing steps).
Video 2. 3D reconstruction of one of Justine’s shell samples
It was a very fruitful week and besides interesting new ideas for scanning our museum specimens, we also met many old and new friends during the week that was as inspiring. ForBio and Transmitting Science did a really great job with setting up this course and I can highly recommend it to anyone who is interested in CT scanning.
Explore the world, read the invertebrate blogs!
-Cessa
Field season’s end
Sletvik field station, October 15th-23rd 2019
- Sletvik field station, Agdenes
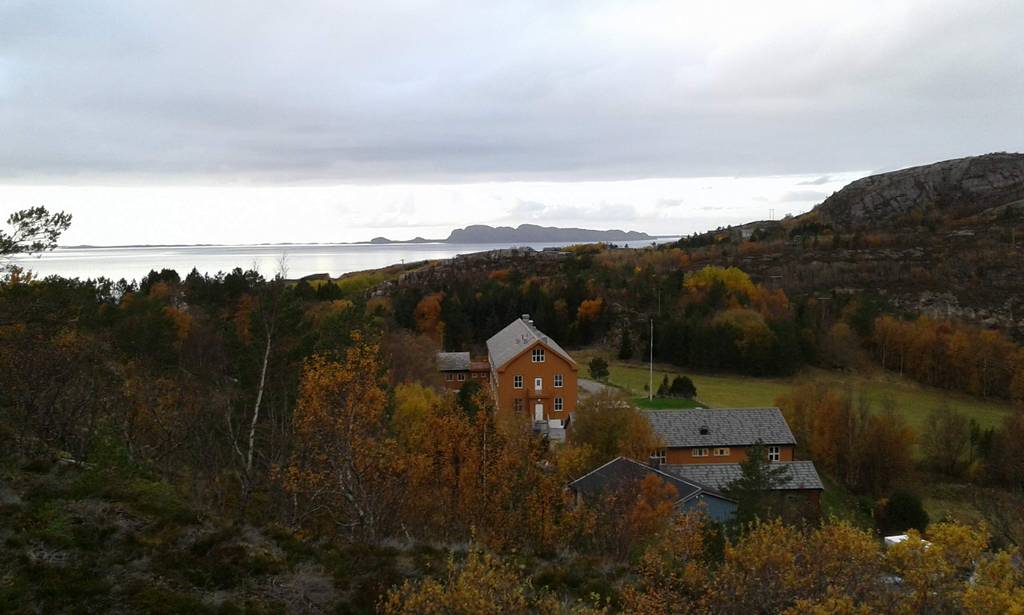
We wanted to make a write-up of the last combined fieldwork/workshop we had in 2019, which was a trip to the marine field station of NTNU, Sletvik in Trøndelag, in late October. From Bergen, Luis (NorHydro), Jon, Tom, and Katrine (Hardbunnsfauna) stuffed a car full of material, microscopes, and drove the ~12 hours up to the field station that we last visited in 2016. Beautiful fall in Trøndelag
Beautiful fall in Trøndelag
There we joined up with Torkild, Aina, Karstein, and Tuva from NTNU university museum, students August and Marte, and Eivind from NIVA. We also had some visitors; Hauk and Stine from Artsdatabanken came by to visit (if you read Norwegian, there’s a feature about it here), and Per Gätzschmann from NTNU UM dropped by for a day to photograph people in the field.
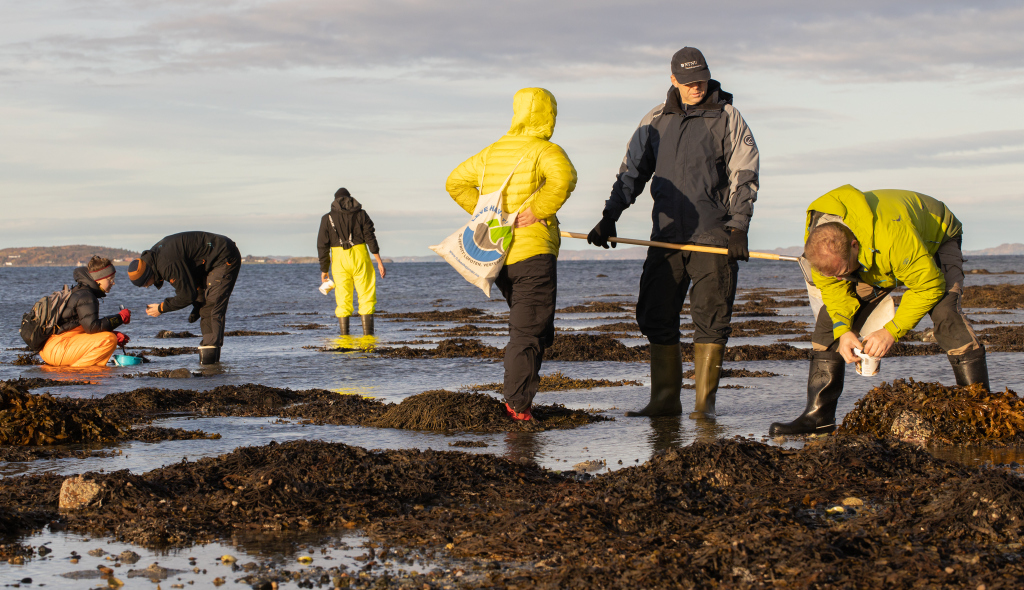
During a productive week the plan was to work through as much as possible of the material that we and our collaborators had collected from Kristiansand in the South to Svalbard in the North. Some of us went out every day to collect fresh material in the field close to the station.The Artsprosjekts #Sneglebuss, Hardbunnsfauna, NorHydro, and PolyPort gathered at Sletvik, and with that also the University museums of Trondheim and Bergen. Of course we were also collecting for the other projects, and the museum collections.
- The littoral zone was teeming with biologists (and some invertebrates). Photo: Per Gätzschmann, NTNU
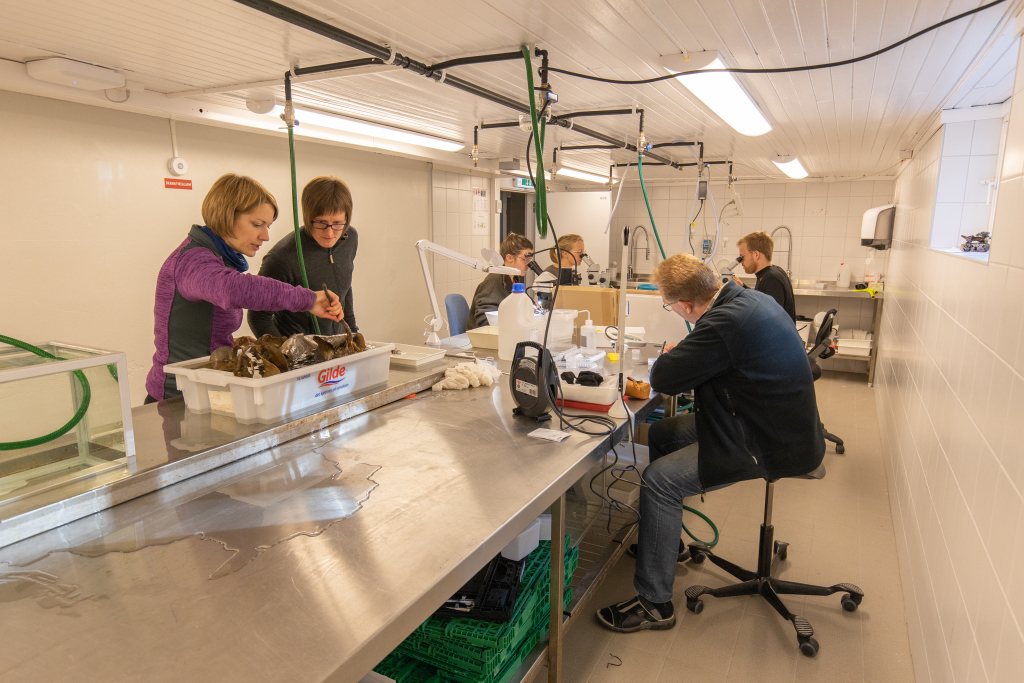
- Busy lab, where we were sorting the fresh samples Photo: Per Gätzschmann, NTNU
- Photo: Per Gätzschmann, NTNU
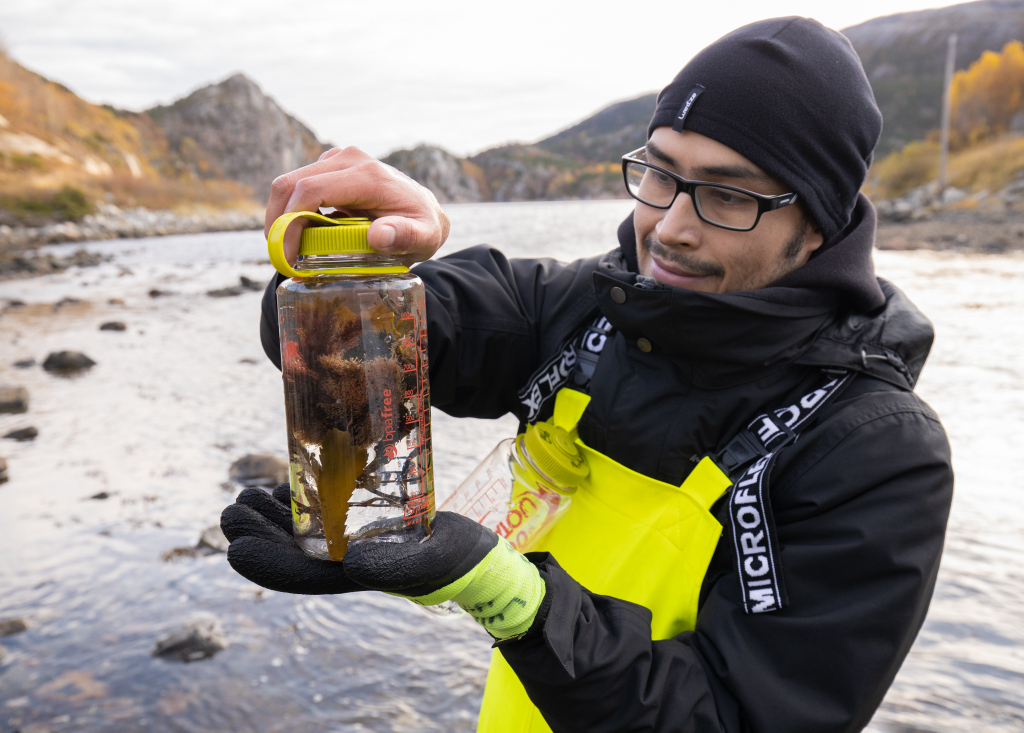
- Luis in the field, showing a catch of hydroids. Photo: Per Gätzschmann, NTNU
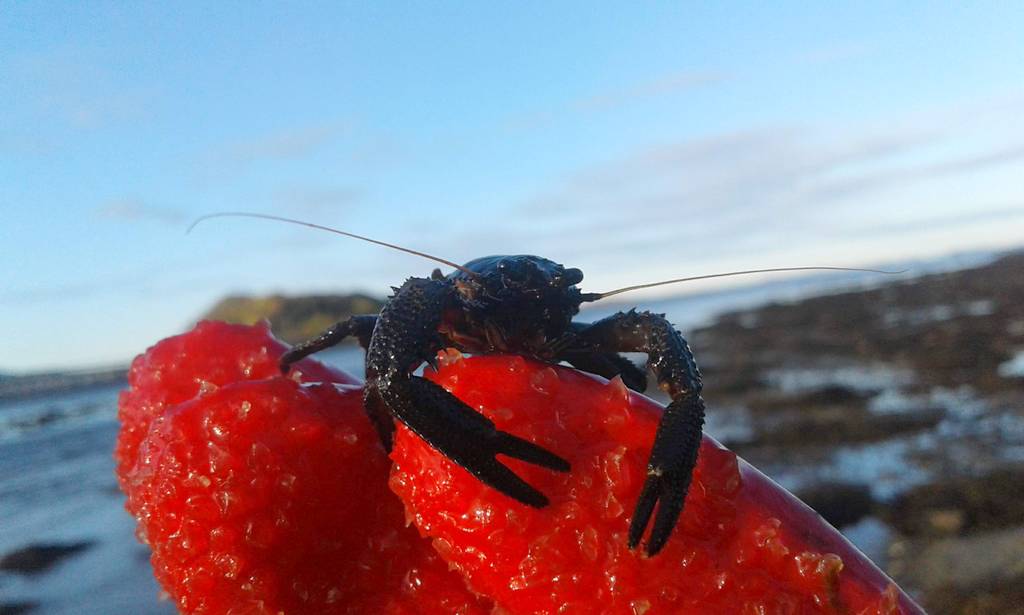
- A squat lobster that would not let go of the red gloves before having its picture taken Photo: K. Kongshavn
- The great Littorina escape; Sletvik edition – we found them all over, as usual. One managed to clog a sink enough to cause a minor flood – that’s when it is good to be in a wet lab.
One of the things Hardbunnsfauna wanted to do whilst in Sletvik was to pick out interesting specimens to submit for DNA barcoding. This means that the animals need to be sorted from the sediment, the specimens identified, and the ones destined to become barcode vouchers must be photographed and tissue sampled, and the data uploaded to the BOLD database. We managed to complete three plates of gastropods, select specimens for one with bivalves, and begin on a plate of echinoderms, as well as sort through and select quite a few crustaceans and ascidians for further study.
- Photolab – trying to convince the amphipods to stand still so we can capture their colours Photo: K. Kongshavn
- Some seaweed to cling to helps
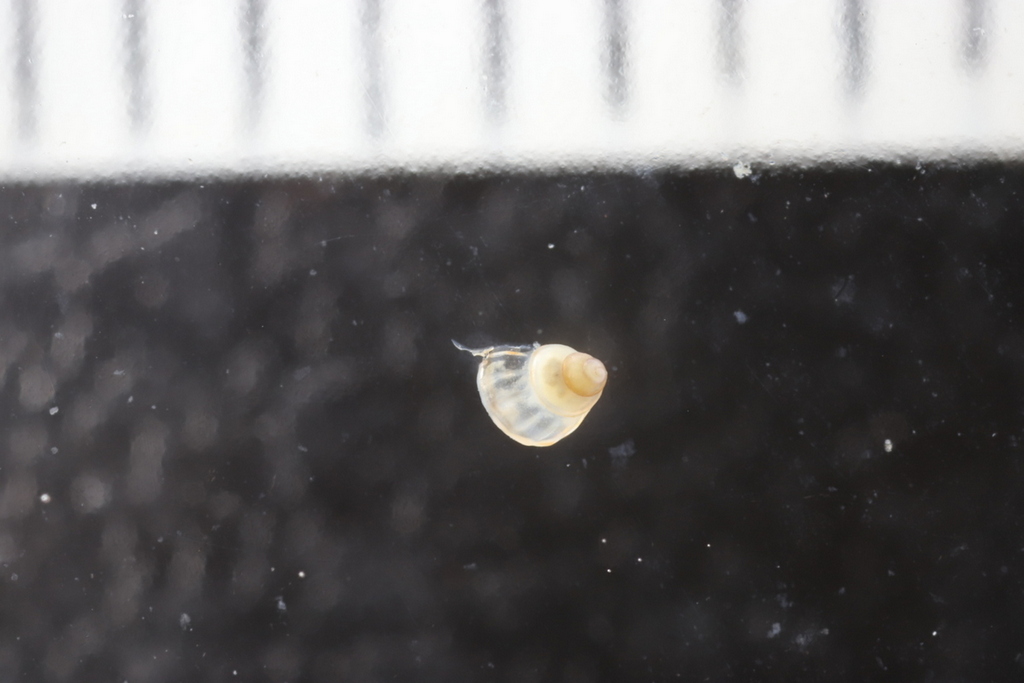
- Many of the gastropods were truly *tiny*, the scale above shows millimeters
- Aina did a fantastic job persuading tissue samples out of the shells of the (mostly minute) snails, and filling the three plates with tissues samples.
- Samples for barcoding
- We also managed ~half a plate of echinoderms, we’ll fill the rest as we go through more material
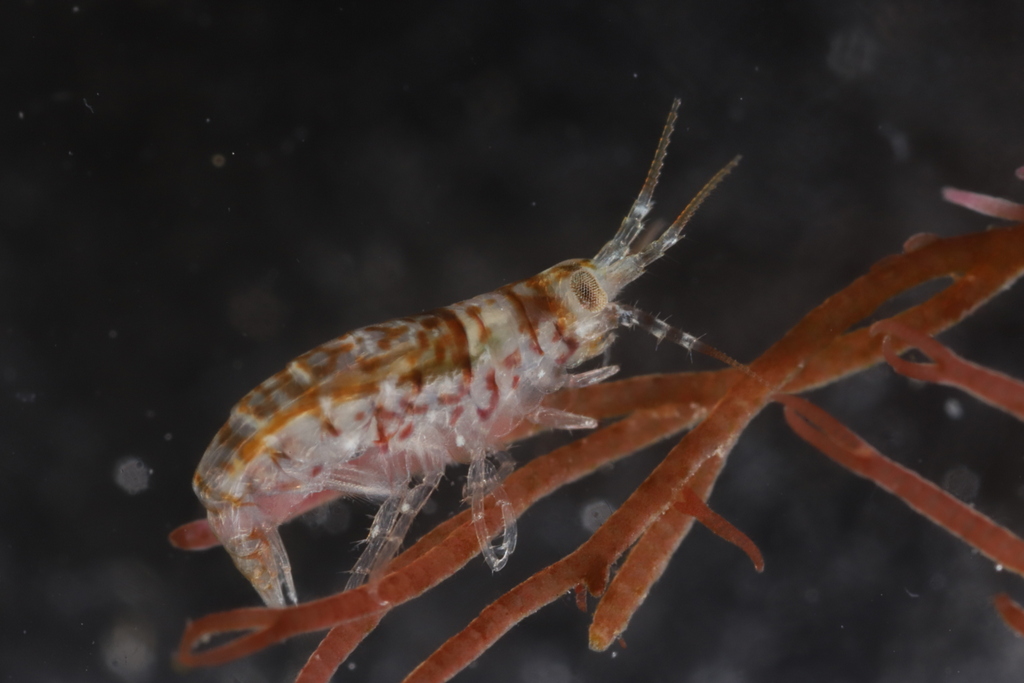
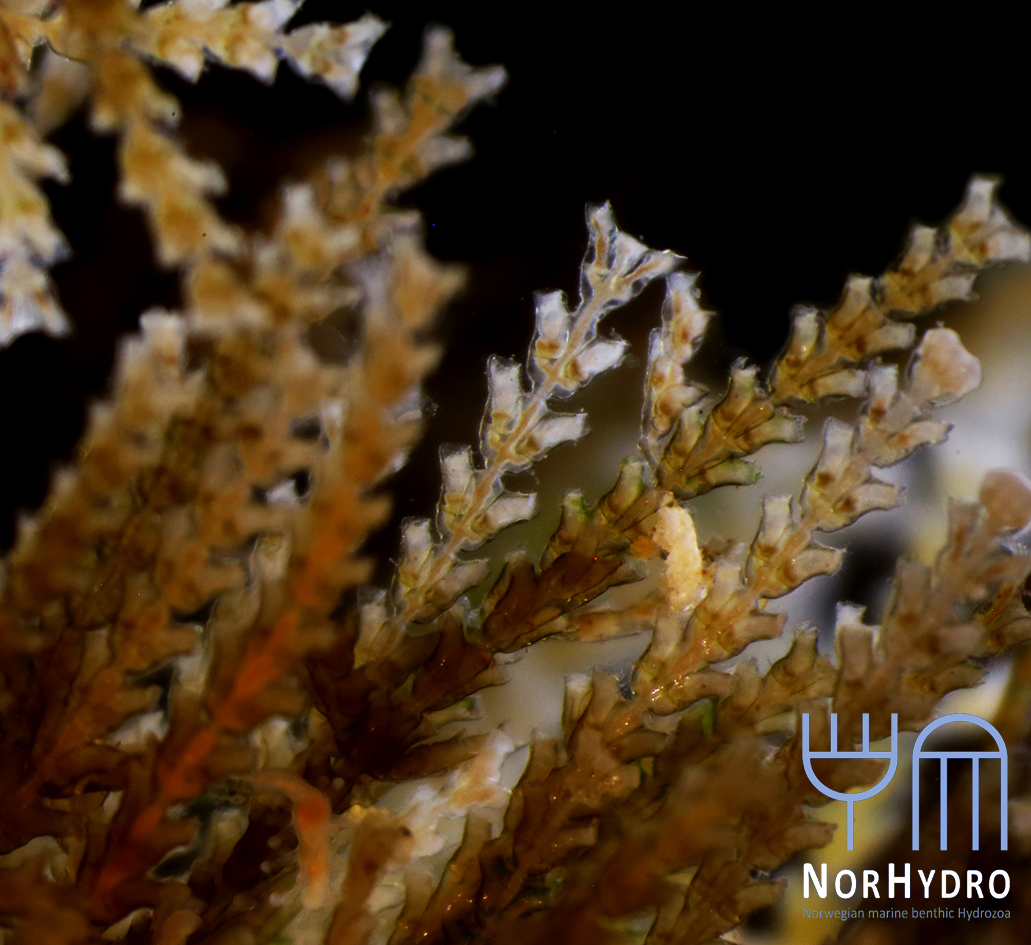
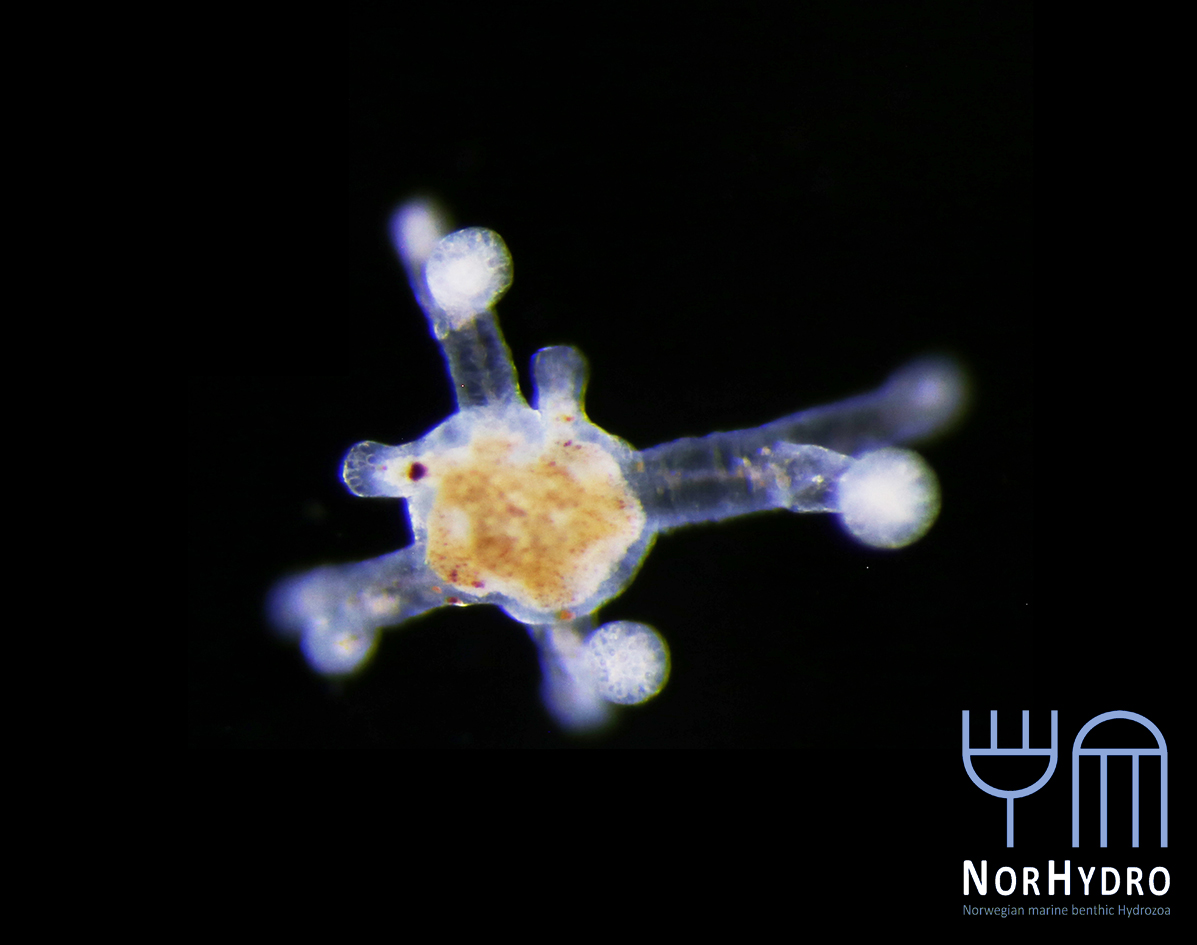
Collecting some fresh material was particularly important for NorHydro because the hydroids from the coasts of Trøndelag have not been thoroughly studied in recent years, and therefore we expected some interesting findings in the six sites we managed to sample. We selected over 40 hydrozoan specimens for DNA barcoding, including some common and widespread hydroids (e.g. Dynamena pumila), some locally abundant species (e.g. Sarsia lovenii) and exceptionally rare taxa, such as the northernmost record ever for a crawling medusa (Eleutheria dichotoma). We also used a small plankton net to catch some of the local hydromedusae, and found many baby jellyfish belonging to genus Clytia swimming around the field station.
- Dynamena pumila. Photo: L. Martell
- Eleutheria dichotoma. Photo: L. Martell
- Clytia sp. Photo: L. Martell

Plan B when the animals (in this case Leuckartiara octona) won’t cooperate and be documented with the fancy camera; bring out the cell phones!
It was a busy week, but combining several projects, bringing together material spanning all of Norway, and working together like this made it extremely productive!
Thank you very much to all the participants, and to all the people who have helped us gather material so far!
-Katrine & Luis