Oslo Natural History Museum 02.12.2019 – 06.12.2019
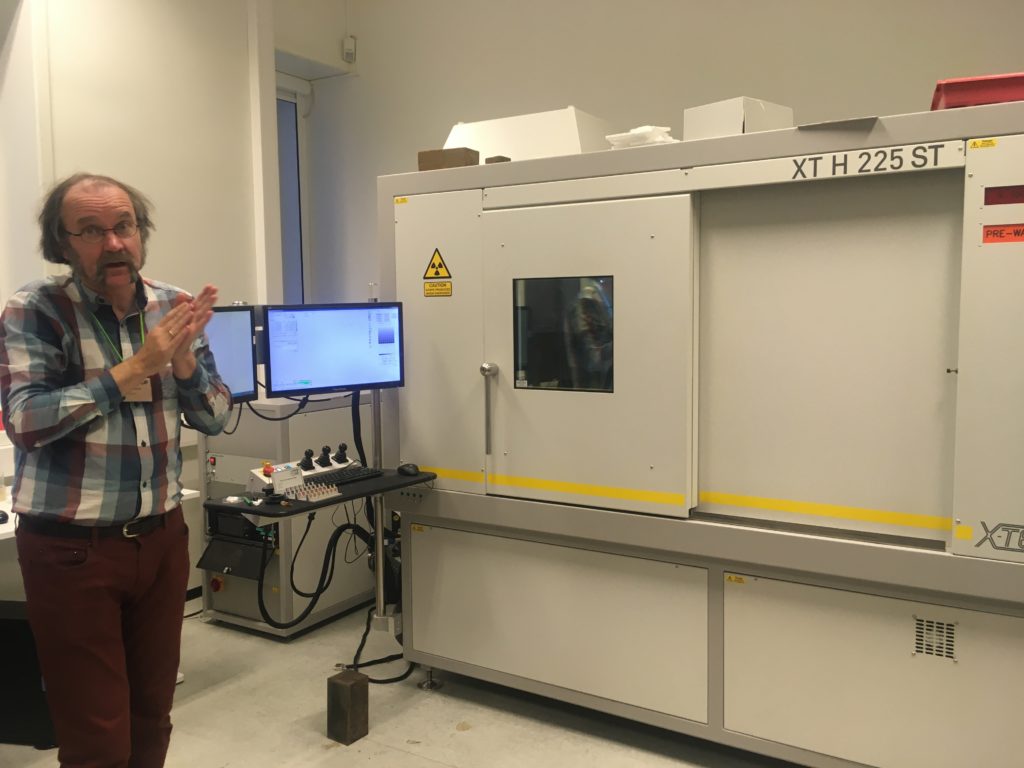
From December 2 till December 6 Justine Siegwald, Manuel Malaquias and I (Cessa) attended a CT scanning course in Oslo at the Natural History Museum.
The course was organized by the Research School in Biosystematics (ForBio) and Transmitting Science.
The three of us all work on mollusks and one of the main reasons to deepen ourselves in CT scanning techniques is because of studying the internal anatomy of these often small and delicate specimens.
Species descriptions are not only based on external morphology and/or DNA barcoding, a lot of the species differences are small details within the internal structures. Like the reproductive organs and digestive system (e.g. radular teeth) of the animals. This is very laborious work, and can be challenging especially with the smaller specimens of only a few millimeters long. Besides some species are difficult to collect and can be a rare collection item of the museum. Once you cut these animals to study them, there is no turning back and the specimen is basically destroyed. Being able to see the insides of the animal without touching it would therefore be ideal. CT scanning comes very close to that and with powerful X-ray we can almost see every detail of the insides of the animals without the need to cut them open
However, how easy as this sounds, CT scanning soft tissue comes with some challenges. Soft tissue means that the x-ray contrast is often very low. Even with modern good x-ray detectors it is still difficult to detect the different internal structures. Therefore, during the course, we were taught how to artificially increase the contrast of the tissue by staining the specimens before mounting them in the CT scan machine.
Coating a specimen with a metal (e.g. gold or platinum) is only useful if you want to see external details of the study object, coating will not help revealing internal structures. For radio contrasting the internal anatomy, you can stain specimens with high density liquids like iodine to enhance the x-ray contrast. So, we went for this option and left our specimens in iodine ethanol solution overnight
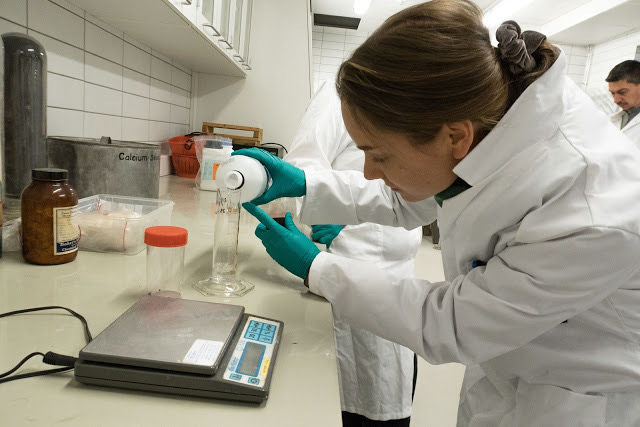
- Preparing the iodine solution
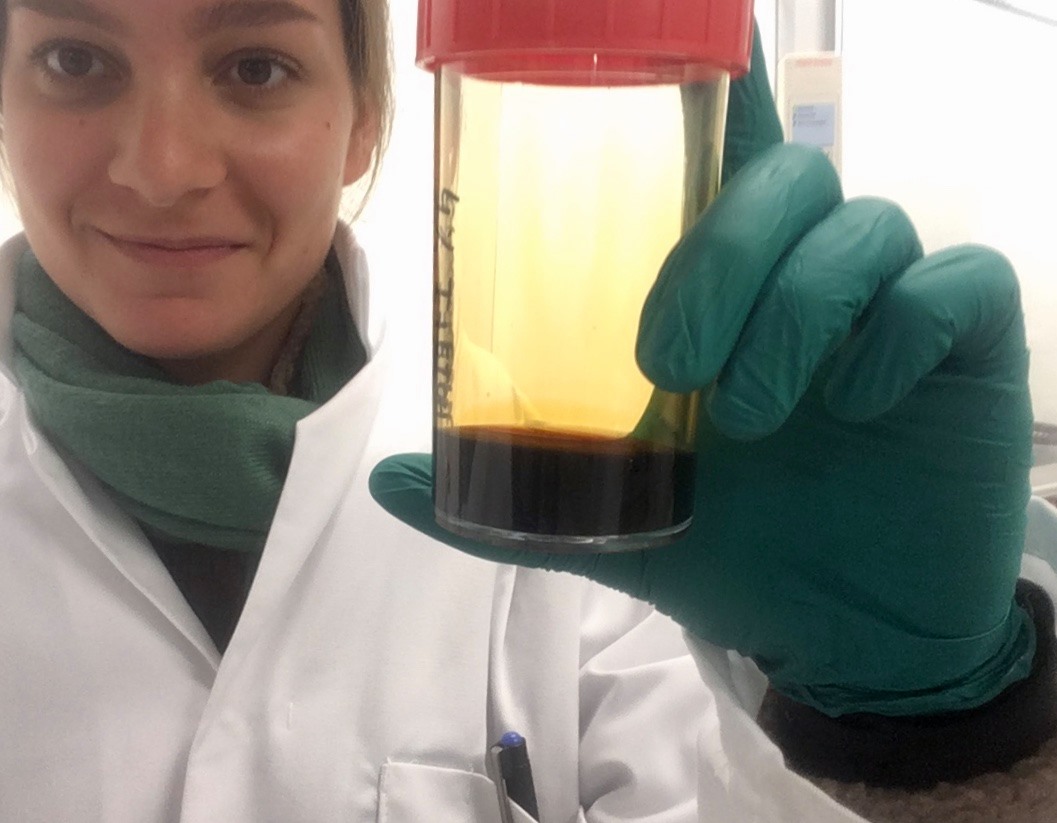
- Sea slug in iodine solution
After the staining process, we needed the wash out any extra iodine before mounting the specimen into the CT scan. A good scan is not simply pressing the button of the machine; there are a bunch of settings that can be adjusted accordingly (e.g. tube voltage, tube current, filament current, spot size, exposure time, magnification, shading correction, etc.). The scan itself can take hours and file sizes of 24GB for a single object are not uncommon, which means you need a powerful computer with decent software to process this information. Part of the course was also about how to visualize and analyze the files of the CT scan with software like Avizo
Video 1. Software Avizo has a user-friendly interface for analyzing all sorts of CT scanning files.
The week gave us some very promising first results (se video below), and also new insights about how to increase the X-ray contrast of our sea slug samples (lots of hours of staining and very short washing steps).
Video 2. 3D reconstruction of one of Justine’s shell samples
It was a very fruitful week and besides interesting new ideas for scanning our museum specimens, we also met many old and new friends during the week that was as inspiring. ForBio and Transmitting Science did a really great job with setting up this course and I can highly recommend it to anyone who is interested in CT scanning.
Explore the world, read the invertebrate blogs!
-Cessa