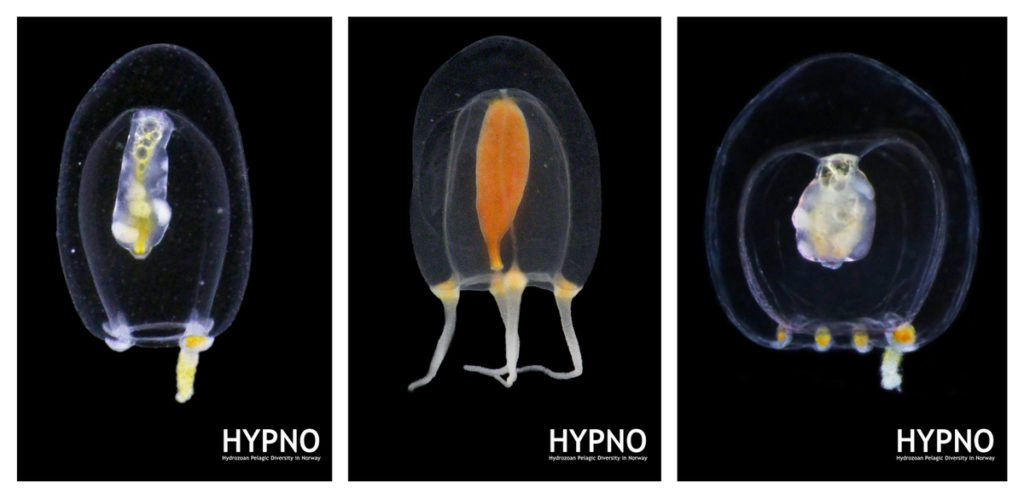
MSc student Ana González visited the collections last month as part of project NorHydro, where she spent some weeks in the lab working with her samples. Here is an account of her experience:
The challenge of identifying benthic hydrozoans
Hydrozoa is a fascinating but poorly understood group of invertebrates, in part because their identification is not always an easy task. I have been studying benthic hydrozoan communities for over a year now, in particular those living in the shallow waters of Mallorca (Spain), and I have realized that the diversity of forms and structures in the group is higher than I had imagined at the beginning of my studies, and their identification is more difficult than I expected. The assemblages of hydrozoans in the Mediterranean are of course very different from the ones that occur in Norway, but something that both communities have in common is that morphological identification of the animals (i.e. telling which species is present based only on the characteristics we can observe) is challenging, which is why one of the aims of my visit to the University Museum of Bergen last December was to learn a different technique (DNA barcoding) that can help me improve the identification of my samples in cases when the morphology of the specimens is not good enough.
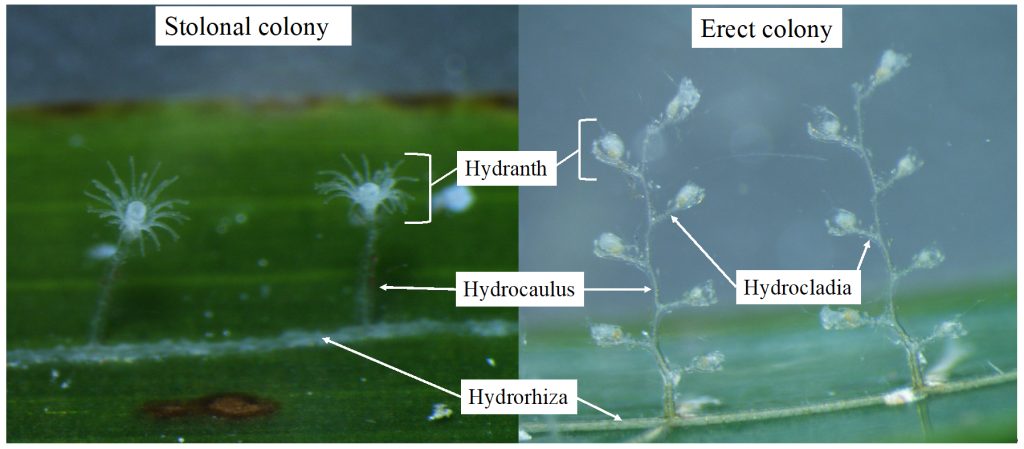

Some of the morphological characters that are used to identify benthic hydrozoans. On the left side a member of Campanulariidae, with a stolonal colony, and on the right side Monotheca obliqua with an erect colony.
DNA barcoding consists in finding a short DNA sequence (the barcode) that is similar for all members of one species but different from all other species. It is a relatively recent tool that –among other things– has helped the scientific community identify specimens that for one reason or the other cannot be identified based on how they look. In some groups, such as many colonial invertebrates, this technique has become a key asset because the colonies are often too young or not reproductive, or the important characters for identifications may be found only in one stage of the life cycle and not in others. For this visit I had the chance to bring all my samples from Mallorca to Bergen and I set to extracting the DNA of selected specimens, amplifying two different barcode genes (COI and 16S), and obtaining clean sequences for them. I discovered that, when it comes to DNA barcoding, every step of the process is important, and being patient and careful is essential.
Getting good results in the DNA lab depends on several factors like not forgetting any step and avoiding contamination as far as possible, but the work does not end there: once you have your sequences they have to be cleaned, quality-checked, and finally compared with others. This means that having a complete and trustworthy database of DNA barcodes is necessary, especially if you want to use the sequence to help you corroborate the identification of a specimen. When done right and with a good database, the DNA barcodes can be useful to detect differences between hydrozoan assemblages growing in different parts of the world or between different substrates and levels of anthropogenic impact, which is what I am doing in my MSc project.
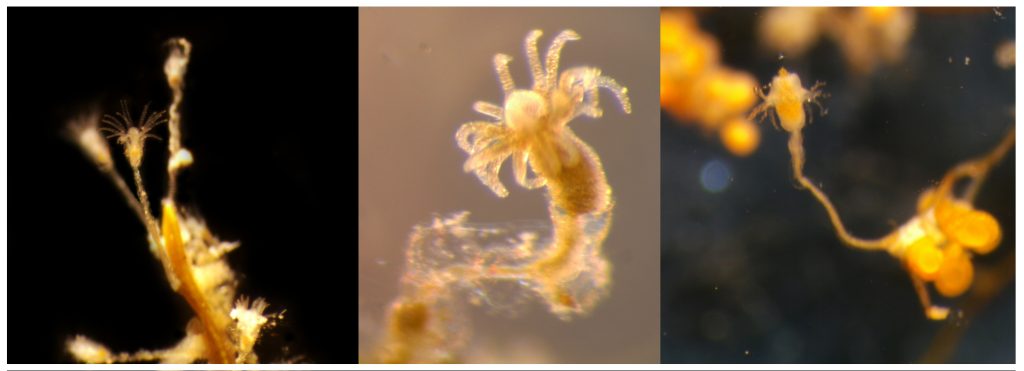
Left: Clytia sp growing on the marine plant Posidonia oceanica. Center: A polyp of Halecium sp, one of the most difficult genera of Hydrozoa to identify based only in morphology, especially when the colony is not reproductive. Right: Eudendrium sp., found in harbours in Mallorca in high abundances.
The analysis of DNA sequences is a powerful tool to compare specimens of distinct populations and in some cases animals that apparently belong to the same species turn out to be completely different (e.g. cryptic species). This is not uncommon for benthic hydrozoans, which have high morphological diversity but also high levels of plasticity, resulting in colonies from different species sometimes being very similar to each other when they grow in similar substrates. As useful as DNA analyses are, however, it is also important to consider their limitations. For example, while the abundance of each species in a given community is important to describe the ecological status of a habitat, estimating abundance is still not always possible from sequence reads in DNA analyses.
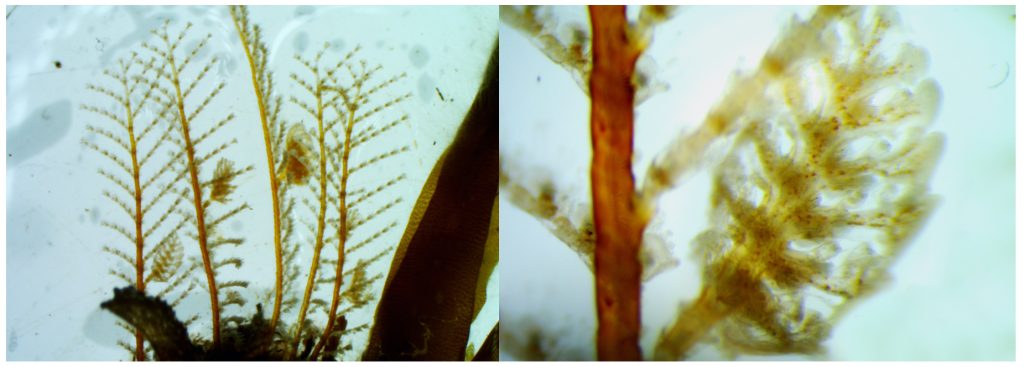

Many cryptic species have been discovered in Aglaopheniidae thanks to the combination of DNA barcoding and morphological analysis
The use of DNA barcodes in my work is not limited to my current project, as I hope my identifications and sequences will help a little bit to improve the databases for future studies of hydrozoan communities in the Mediterranean Sea, and maybe even allow other researchers to compare their samples with the species found on other parts of the world. I think that looking closely at each specimen is the best way to truly know variation, so both morphology observations and DNA analyses should be combined to obtain good estimates of the diversity of a taxon in any locality. For example, whenever the DNA analyses reveal differences in two clades that were thought to be the same species, it is time to search for new taxonomic characters that we might have missed before, and for that reason it is also important to have a good knowledge of the morphology of each species. Both morphological and DNA-based identifications have limitations and advantages so, if you have the opportunity to use both, why choose only one?
Ana